Design and evaluation of 1,7-naphthyridones as novel KDM5 inhibitors.
Labadie, S.S., Dragovich, P.S., Cummings, R.T., Deshmukh, G., Gustafson, A., Han, N., Harmange, J.C., Kiefer, J.R., Li, Y., Liang, J., Liederer, B.M., Liu, Y., Manieri, W., Mao, W., Murray, L., Ortwine, D.F., Trojer, P., VanderPorten, E., Vinogradova, M., Wen, L.(2016) Bioorg Med Chem Lett 26: 4492-4496
- PubMed: 27499454 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2016.07.070
- Primary Citation Related Structures:
5K4L - PubMed Abstract:
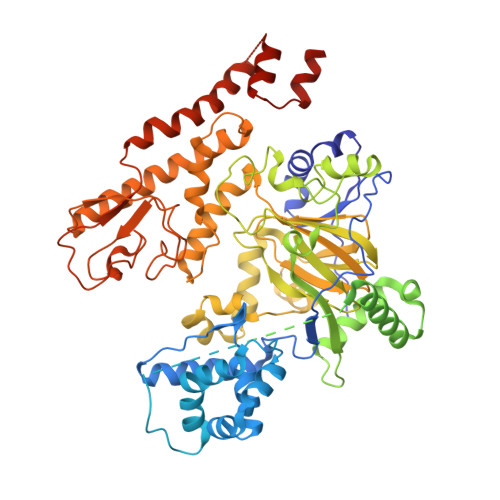
Features from a high throughput screening (HTS) hit and a previously reported scaffold were combined to generate 1,7-naphthyridones as novel KDM5 enzyme inhibitors with nanomolar potencies. These molecules exhibited high selectivity over the related KDM4C and KDM2B isoforms. An X-ray co-crystal structure of a representative molecule bound to KDM5A showed that these inhibitors are competitive with the co-substrate (2-oxoglutarate or 2-OG).
- Genentech Inc., 1 DNA Way, South San Francisco, CA 94080, USA.
Organizational Affiliation: