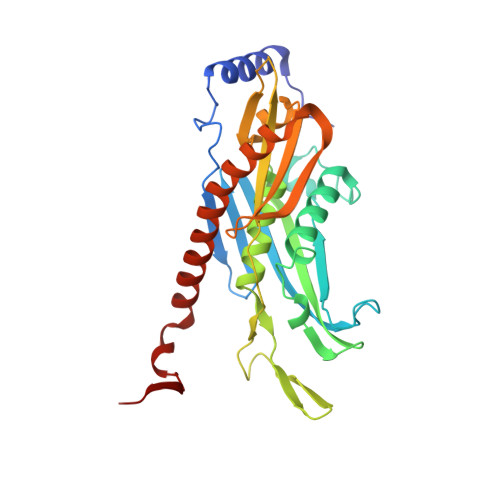
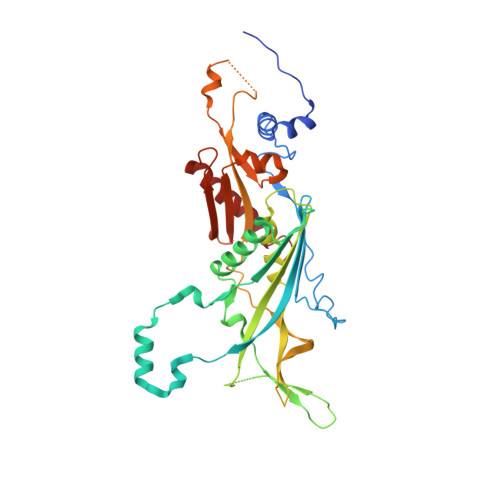
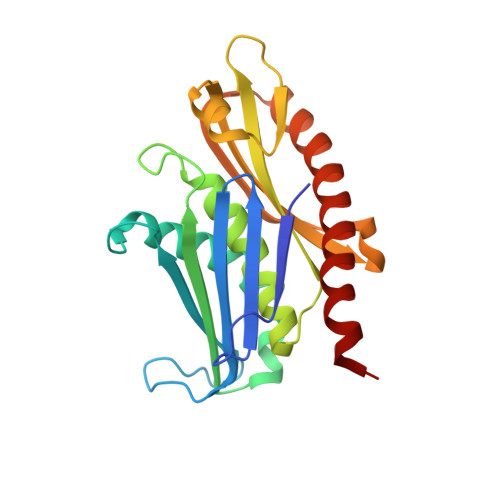
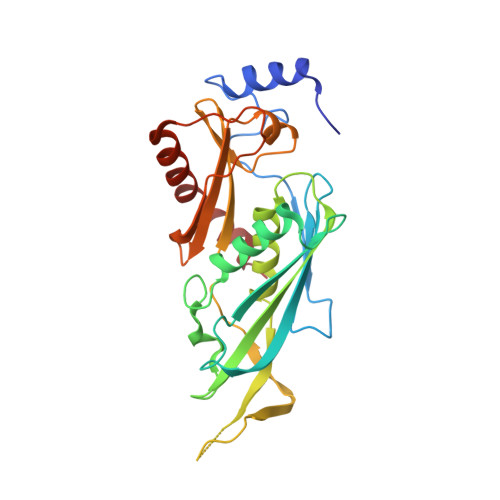
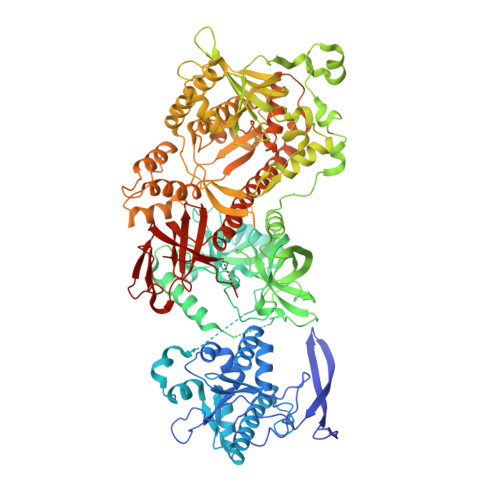
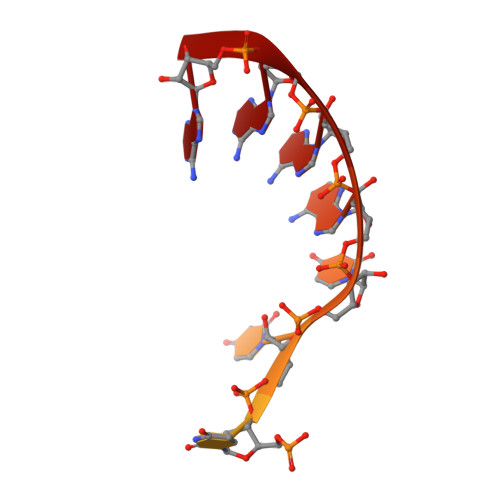
Nuclear RNA Exosome at 3.1 angstrom Reveals Substrate Specificities, RNA Paths, and Allosteric Inhibition of Rrp44/Dis3.
Zinder, J.C., Wasmuth, E.V., Lima, C.D.(2016) Mol Cell 64: 734-745
- PubMed: 27818140 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2016.09.038
- Primary Citation Related Structures:
5K36 - PubMed Abstract:
The eukaryotic RNA exosome is an essential and conserved 3'-to-5' exoribonuclease complex that degrades or processes nearly every class of cellular RNA. The nuclear RNA exosome includes a 9-subunit non-catalytic core that binds Rrp44 (Dis3) and Rrp6 subunits to modulate their processive and distributive 3'-to-5' exoribonuclease activities, respectively. Here we utilize an engineered RNA with two 3' ends to obtain a crystal structure of an 11-subunit nuclear exosome bound to RNA at 3.1 Å. The structure reveals an extended RNA path to Rrp6 that penetrates into the non-catalytic core; contacts between the non-catalytic core and Rrp44, which inhibit exoribonuclease activity; and features of the Rrp44 exoribonuclease site that support its ability to degrade 3' phosphate RNA substrates. Using reconstituted exosome complexes, we show that 3' phosphate RNA is not a substrate for Rrp6 but is readily degraded by Rrp44 in the nuclear exosome.
- Tri-Institutional Training Program in Chemical Biology, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA; Structural Biology Program, Sloan Kettering Institute, 1275 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: