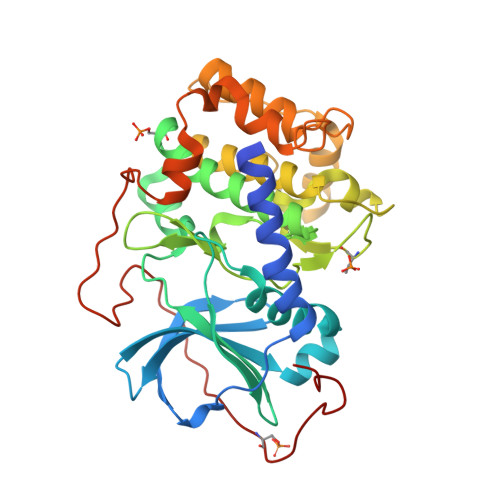
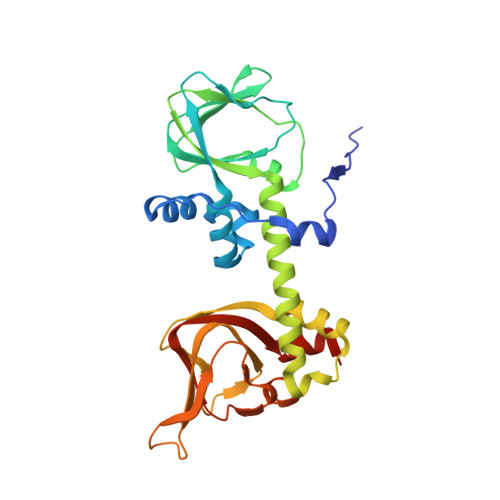
Structure of a PKA RI alpha Recurrent Acrodysostosis Mutant Explains Defective cAMP-Dependent Activation.
Bruystens, J.G., Wu, J., Fortezzo, A., Del Rio, J., Nielsen, C., Blumenthal, D.K., Rock, R., Stefan, E., Taylor, S.S.(2016) J Mol Biology 428: 4890-4904
- PubMed: 27825928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2016.10.033
- Primary Citation Related Structures:
5JR7 - PubMed Abstract:
Most disease-related mutations that impair cAMP protein kinase A (PKA) signaling are present within the regulatory (R) PKA RI alpha-subunit (RIα). Although mutations in the PRKAR1A gene are linked to Carney complex (CNC) disease and, more recently, to acrodysostosis-1 (ACRDYS1), the two diseases show contrasting phenotypes. While CNC mutations cause increased PKA activity, ACRDYS1 mutations result in decreased PKA activity and cAMP resistant holoenzymes. Mapping the ACRDYS1 disease mutations reveals their localization to the second of two tandem cAMP-binding (CNB) domains (CNB-B), and here, we characterize a recurrent deletion mutant where the last 14 residues are missing. The crystal structure of a monomeric form of this mutant (RIα92-365) bound to the catalytic (C)-subunit reveals the dysfunctional regions of the RIα subunit. Beyond the missing residues, the entire capping motif is disordered (residues 357-379) and explains the disrupted cAMP binding. Moreover, the effects of the mutation extend far beyond the CNB-B domain and include the active site and N-lobe of the C-subunit, which is in a partially open conformation with the C-tail disordered. A key residue that contributes to this crosstalk, D267, is altered in our structure, and we confirmed its functional importance by mutagenesis. In particular, the D267 interaction with Arg241, a residue shown earlier to be important for allosteric regulation, is disrupted, thereby strengthening the interaction of D267 with the C-subunit residue Arg194 at the R:C interface. We see here how the switch between active (cAMP-bound) and inactive (holoenzyme) conformations is perturbed and how the dynamically controlled crosstalk between the helical domains of the two CNB domains is necessary for the functional regulation of PKA activity.
- Department of Chemistry and Biochemistry, University of California at San Diego, La Jolla, CA 92093, USA.
Organizational Affiliation: