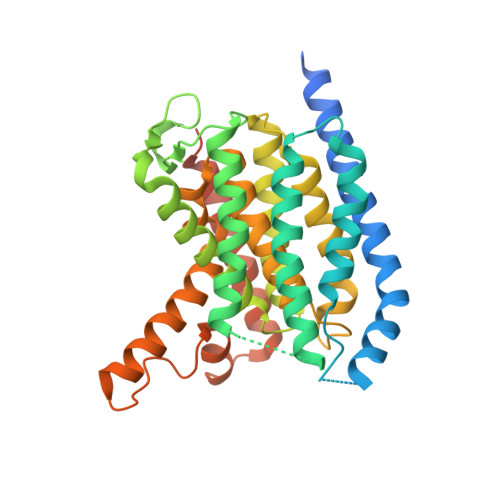
MraY-antibiotic complex reveals details of tunicamycin mode of action.
Hakulinen, J.K., Hering, J., Branden, G., Chen, H., Snijder, A., Ek, M., Johansson, P.(2017) Nat Chem Biol 13: 265-267
- PubMed: 28068312 Search on PubMed
- DOI: https://doi.org/10.1038/nchembio.2270
- Primary Citation Related Structures:
5JNQ - PubMed Abstract:
The rapid increase of antibiotic resistance has created an urgent need to develop novel antimicrobial agents. Here we describe the crystal structure of the promising bacterial target phospho-N-acetylmuramoyl-pentapeptide translocase (MraY) in complex with the nucleoside antibiotic tunicamycin. The structure not only reveals the mode of action of several related natural-product antibiotics but also gives an indication on the binding mode of the MraY UDP-MurNAc-pentapeptide and undecaprenyl-phosphate substrates.
- Discovery Sciences, Innovative Medicines and Early Development Biotech Unit, AstraZeneca R&D Gothenburg, Gothenburg, Sweden.
Organizational Affiliation: