Discovery of acyl guanidine tryptophan hydroxylase-1 inhibitors.
Goldberg, D.R., De Lombaert, S., Aiello, R., Bourassa, P., Barucci, N., Zhang, Q., Paralkar, V., Stein, A.J., Valentine, J., Zavadoski, W.(2016) Bioorg Med Chem Lett 26: 2855-2860
- PubMed: 27146606 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2016.04.057
- Primary Citation Related Structures:
5J6D - PubMed Abstract:
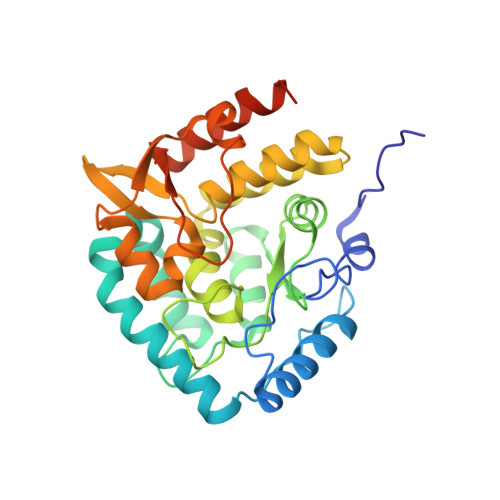
An increasing number of diseases have been linked to a dysfunctional peripheral serotonin system. Given that tryptophan hydroxylase 1 (TPH1) is the rate limiting enzyme in the biosynthesis off serotonin, it represents an attractive target to regulate peripheral serotonin. Following up to our first disclosure, we report a new chemotype of TPH1 inhibitors where-by the more common central planar heterocycle has been replaced with an open-chain, acyl guanidine surrogate. Through our work, we found that compounds of this nature provide highly potent TPH1 inhibitors with favorable physicochemical properties that were effective in reducing murine intestinal 5-HT in vivo. Furthermore, we obtained a high resolution (1.90Å) X-ray structure crystal structure of one of these inhibitors (compound 51) that elucidated the active conformation along with revealing a dimeric form of TPH1 for the first time.
- Karos Pharmaceuticals, 401 Winchester Ave., 5 Science Park, New Haven, CT 06511, United States. Electronic address: dgoldberg@kineta.us.
Organizational Affiliation: