A Novel Class Of Virulence Factors In Penicillium Marneffei And Aspergillus Fumigatus Enhances Intracellular Survival In Monocytes By Arachidonic Acid Binding
Zhang, H., Hao, Q., Lam, W.H.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Uncharacterized protein | 164 | Aspergillus fumigatus Af293 | Mutation(s): 0 Gene Names: AFUA_2G17630 | 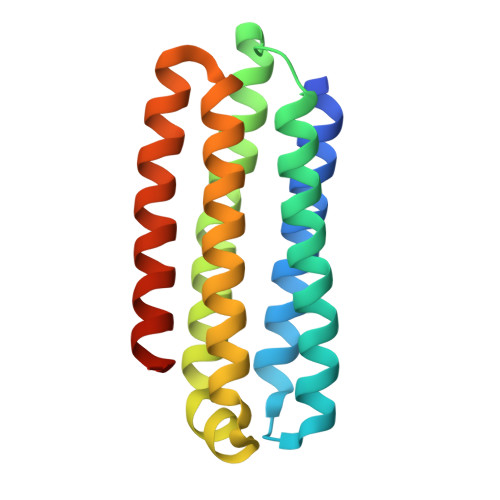 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q4WZA5 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PLM Download:Ideal Coordinates CCD File | C [auth A] | PALMITIC ACID C16 H32 O2 IPCSVZSSVZVIGE-UHFFFAOYSA-N |  | ||
| MAN Download:Ideal Coordinates CCD File | B [auth A] | alpha-D-mannopyranose C6 H12 O6 WQZGKKKJIJFFOK-PQMKYFCFSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 124.836 | α = 90 |
| b = 124.836 | β = 90 |
| c = 124.836 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| PHASER | phasing |
| REFMAC | refinement |