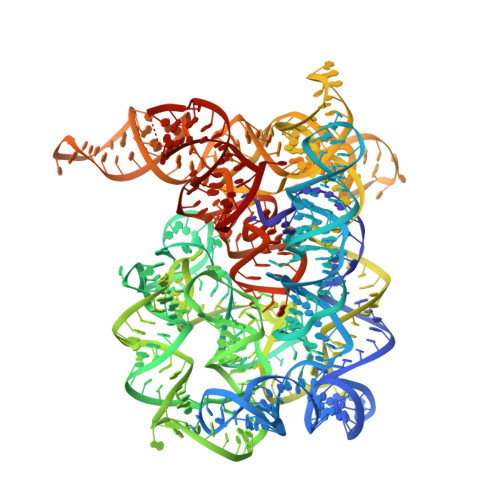
Crystal structures of a group II intron lariat primed for reverse splicing.
Costa, M., Walbott, H., Monachello, D., Westhof, E., Michel, F.(2016) Science 354
- PubMed: 27934709 Search on PubMed
- DOI: https://doi.org/10.1126/science.aaf9258
- Primary Citation Related Structures:
5J01, 5J02 - PubMed Abstract:
The 2'-5' branch of nuclear premessenger introns is believed to have been inherited from self-splicing group II introns, which are retrotransposons of bacterial origin. Our crystal structures at 3.4 and 3.5 angstrom of an excised group II intron in branched ("lariat") form show that the 2'-5' branch organizes a network of active-site tertiary interactions that position the intron terminal 3'-hydroxyl group into a configuration poised to initiate reverse splicing, the first step in retrotransposition. Moreover, the branchpoint and flanking helices must undergo a base-pairing switch after branch formation. A group II-based model of the active site of the nuclear splicing machinery (the spliceosome) is proposed. The crucial role of the lariat conformation in active-site assembly and catalysis explains its prevalence in modern splicing.
- Group II introns as ribozymes and retrotransposons, Institute for Integrative Biology of the Cell (I2BC), UMR 9198 CNRS, Commissariat à l'Energie Atomique et aux Energies Alternatives (CEA), University Paris-Sud, University Paris-Saclay, 1 Avenue de la Terrasse, Bâtiment 26, 91198 Gif-sur-Yvette cedex, France. maria.costa@i2bc.paris-saclay.fr.
Organizational Affiliation: