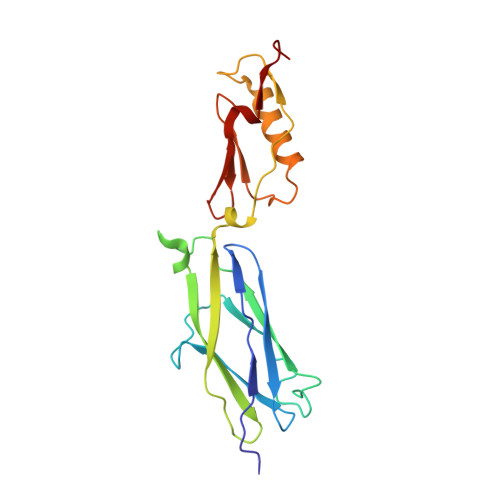
Structures of the Streptococcus sanguinis SrpA Binding Region with Human Sialoglycans Suggest Features of the Physiological Ligand.
Loukachevitch, L.V., Bensing, B.A., Yu, H., Zeng, J., Chen, X., Sullam, P.M., Iverson, T.M.(2016) Biochemistry
- PubMed: 27685666 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00704
- Primary Citation Related Structures:
5IIY, 5IJ1, 5IJ2, 5IJ3, 5KIQ - PubMed Abstract:
Streptococcus sanguinis is a leading cause of bacterial infective endocarditis, a life-threatening infection of heart valves. S. sanguinis binds to human platelets with high avidity, and this adherence is likely to enhance virulence. Previous studies suggest that a serine-rich repeat adhesin termed SrpA mediates the binding of S. sanguinis to human platelets via its interaction with sialoglycans on the receptor GPIbα. However, in vitro binding assays with SrpA and defined sialoglycans failed to identify specific high-affinity ligands. To improve our understanding of the interaction between SrpA and human platelets, we determined cocrystal structures of the SrpA sialoglycan binding region (SrpA BR ) with five low-affinity ligands: three sialylated trisaccharides (sialyl-T antigen, 3'-sialyllactose, and 3'-sialyl-N-acetyllactosamine), a sialylated tetrasaccharide (sialyl-Lewis X ), and a sialyl galactose disaccharide component common to these sialoglyans. We then combined structural analysis with mutagenesis to further determine whether our observed interactions between SrpA BR and glycans are important for binding to platelets and to better map the binding site for the physiological receptor. We found that the sialoglycan binding site of SrpA BR is significantly larger than the sialoglycans cocrystallized in this study, which suggests that binding of SrpA to platelets either is multivalent or occurs via a larger, disialylated glycan.
- Division of Infectious Diseases, Veterans Affairs Medical Center, University of California, San Francisco and Northern California Institute for Research and Education , San Francisco, California 94121, United States.
Organizational Affiliation: