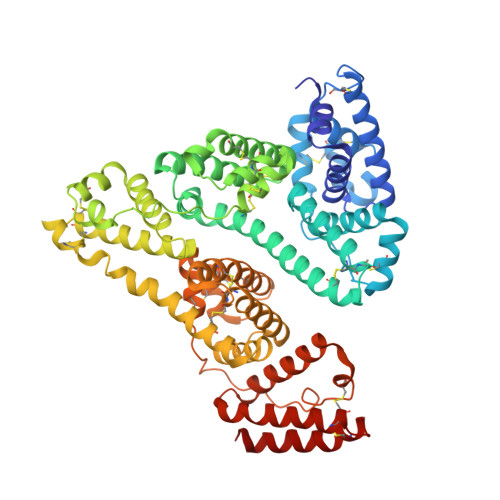
X-ray Structure Analysis of Indazolium trans-[Tetrachlorobis(1H-indazole)ruthenate(III)] (KP1019) Bound to Human Serum Albumin Reveals Two Ruthenium Binding Sites and Provides Insights into the Drug Binding Mechanism.
Bijelic, A., Theiner, S., Keppler, B.K., Rompel, A.(2016) J Med Chem 59: 5894-5903
- PubMed: 27196130 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00600
- Primary Citation Related Structures:
5IFO - PubMed Abstract:
Ruthenium(III) complexes are promising candidates for anticancer drugs, especially the clinically studied indazolium trans-[tetrachlorobis(1H-indazole)ruthenate(III)] (KP1019) and its analogue sodium trans-[tetrachlorobis(1H-indazole)ruthenate(III)] (NKP-1339). Several studies have emphasized the likely role of human serum proteins in the transportation and accumulation of ruthenium(III) complexes in tumors. Therefore, the interaction between KP1019 and human serum albumin was investigated by means of X-ray crystallography and inductively coupled plasma mass spectrometry (ICP-MS). The structural data unambiguously reveal the binding of two ruthenium atoms to histidine residues 146 and 242, which are both located within well-known hydrophobic binding pockets of albumin. The ruthenium centers are octahedrally coordinated by solvent molecules revealing the dissociation of both indazole ligands from the ruthenium-based drug. However, a binding mechanism is proposed indicating the importance of the indazole ligands for binding site recognition and thus their indispensable role for the binding of KP1019.
- Fakultät für Chemie, Institut für Biophysikalische Chemie, Universität Wien , Althanstraße 14, 1090 Wien, Austria.
Organizational Affiliation: