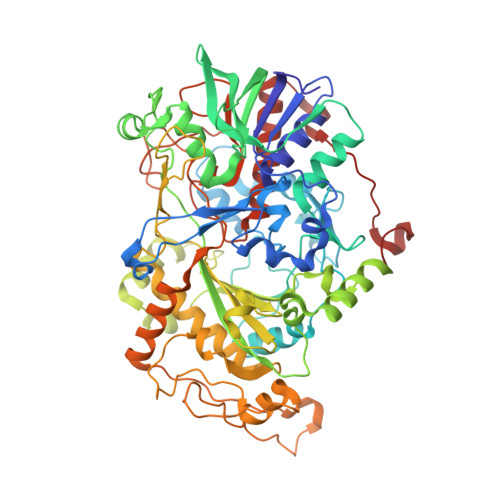
Crystal Structure of Alcohol Oxidase from Pichia pastoris.
Koch, C., Neumann, P., Valerius, O., Feussner, I., Ficner, R.(2016) PLoS One 11: e0149846-e0149846
- PubMed: 26905908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0149846
- Primary Citation Related Structures:
5HSA - PubMed Abstract:
FAD-dependent alcohol oxidases (AOX) are key enzymes of methylotrophic organisms that can utilize lower primary alcohols as sole source of carbon and energy. Here we report the crystal structure analysis of the methanol oxidase AOX1 from Pichia pastoris. The crystallographic phase problem was solved by means of Molecular Replacement in combination with initial structure rebuilding using Rosetta model completion and relaxation against an averaged electron density map. The subunit arrangement of the homo-octameric AOX1 differs from that of octameric vanillyl alcohol oxidase and other dimeric or tetrameric alcohol oxidases, due to the insertion of two large protruding loop regions and an additional C-terminal extension in AOX1. In comparison to other alcohol oxidases, the active site cavity of AOX1 is significantly reduced in size, which could explain the observed preference for methanol as substrate. All AOX1 subunits of the structure reported here harbor a modified flavin adenine dinucleotide, which contains an arabityl chain instead of a ribityl chain attached to the isoalloxazine ring.
- Department of Plant Biochemistry, Albrecht-von-Haller-Institute, Georg-August-University Goettingen, Justus-von-Liebig-Weg 11, 37077, Goettingen, Germany.
Organizational Affiliation: