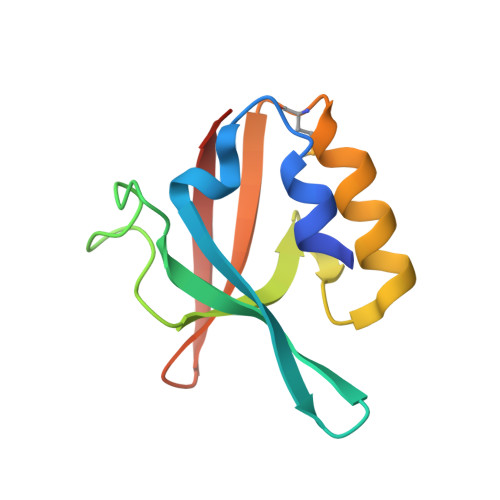
Crystal structure of PDZ domain of pullulanase C protein of type II secretion system from Klebsiella pneumoniae in complex with fatty acid
Filippova, E.V., Minasov, G., Shuvalova, L., Kiryukhina, O., Dubrovska, I., Grimshaw, S., Kwon, K., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.