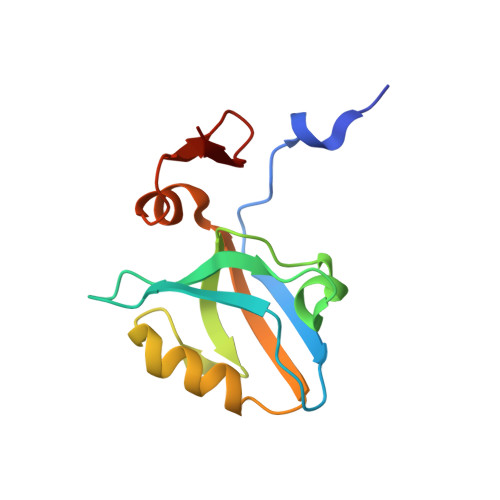
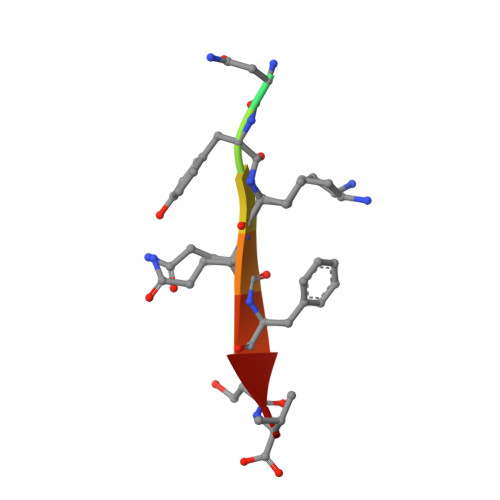
Origins of Allostery and Evolvability in Proteins: A Case Study.
Raman, A.S., White, K.I., Ranganathan, R.(2016) Cell 166: 468-480
- PubMed: 27321669 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2016.05.047
- Primary Citation Related Structures:
5HEB, 5HED, 5HET, 5HEY, 5HF1, 5HFB, 5HFC, 5HFF - PubMed Abstract:
Proteins display the capacity for adaptation to new functions, a property critical for evolvability. But what structural principles underlie the capacity for adaptation? Here, we show that adaptation to a physiologically distinct class of ligand specificity in a PSD95, DLG1, ZO-1 (PDZ) domain preferentially occurs through class-bridging intermediate mutations located distant from the ligand-binding site. These mutations provide a functional link between ligand classes and demonstrate the principle of "conditional neutrality" in mediating evolutionary adaptation. Structures show that class-bridging mutations work allosterically to open up conformational plasticity at the active site, permitting novel functions while retaining existing function. More generally, the class-bridging phenotype arises from mutations in an evolutionarily conserved network of coevolving amino acids in the PDZ family (the sector) that connects the active site to distant surface sites. These findings introduce the concept that allostery in proteins could have its origins not in protein function but in the capacity to adapt.
- Green Center for Systems Biology, University of Texas Southwestern Medical Center, 6001 Forest Park Road, Dallas, TX 75390, USA.
Organizational Affiliation: