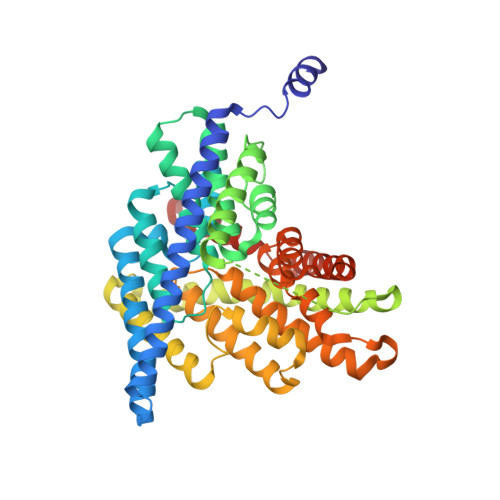
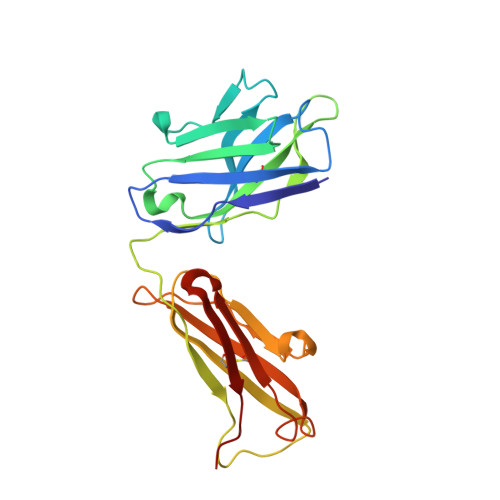
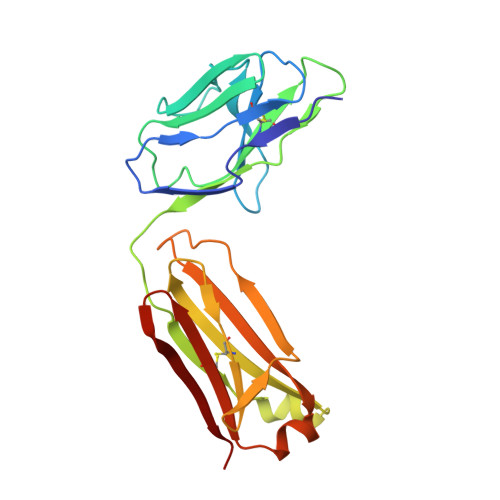
Conformational change in CLC transporters: beyond the rotation of Gluex
Khantwal, C.M., Abraham, S.J., Han, W., Jiang, T., Chavan, T.S., Cheng, R.C., Elvington, S.M., Liu, C.W., Mathews, I.I., Stein, R.A., Mchaourab, H.S., Tajkhorshid, E., Maduke, M.To be published.