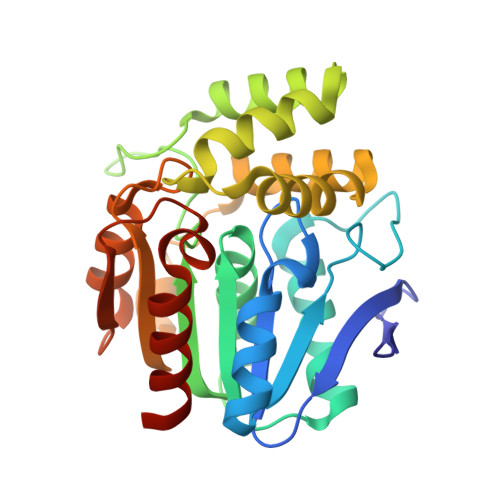
Crystal Structure and Functional Characterization of an Esterase (EaEST) from Exiguobacterium antarcticum.
Lee, C.W., Kwon, S., Park, S.H., Kim, B.Y., Yoo, W., Ryu, B.H., Kim, H.W., Shin, S.C., Kim, S., Park, H., Kim, T.D., Lee, J.H.(2017) PLoS One 12: e0169540-e0169540
- PubMed: 28125606 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0169540
- Primary Citation Related Structures:
5H3H - PubMed Abstract:
A novel microbial esterase, EaEST, from a psychrophilic bacterium Exiguobacterium antarcticum B7, was identified and characterized. To our knowledge, this is the first report describing structural analysis and biochemical characterization of an esterase isolated from the genus Exiguobacterium. Crystal structure of EaEST, determined at a resolution of 1.9 Å, showed that the enzyme has a canonical α/β hydrolase fold with an α-helical cap domain and a catalytic triad consisting of Ser96, Asp220, and His248. Interestingly, the active site of the structure of EaEST is occupied by a peracetate molecule, which is the product of perhydrolysis of acetate. This result suggests that EaEST may have perhydrolase activity. The activity assay showed that EaEST has significant perhydrolase and esterase activity with respect to short-chain p-nitrophenyl esters (≤C8), naphthyl derivatives, phenyl acetate, and glyceryl tributyrate. However, the S96A single mutant had low esterase and perhydrolase activity. Moreover, the L27A mutant showed low levels of protein expression and solubility as well as preference for different substrates. On conducting an enantioselectivity analysis using R- and S-methyl-3-hydroxy-2-methylpropionate, a preference for R-enantiomers was observed. Surprisingly, immobilized EaEST was found to not only retain 200% of its initial activity after incubation for 1 h at 80°C, but also retained more than 60% of its initial activity after 20 cycles of reutilization. This research will serve as basis for future engineering of this esterase for biotechnological and industrial applications.
- Unit of Polar Genomics, Korea Polar Research Institute, Incheon, Republic of Korea.
Organizational Affiliation: