The Adhesion of Lactobacillus salivarius REN to a Human Intestinal Epithelial Cell Line Requires S-layer Proteins
Wang, R., Jiang, L., Zhang, M., Zhao, L., Hao, Y., Guo, H., Sang, Y., Zhang, H., Ren, F.(2017) Sci Rep 7: 44029-44029
- PubMed: 28281568 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep44029
- Primary Citation Related Structures:
5GT1 - PubMed Abstract:
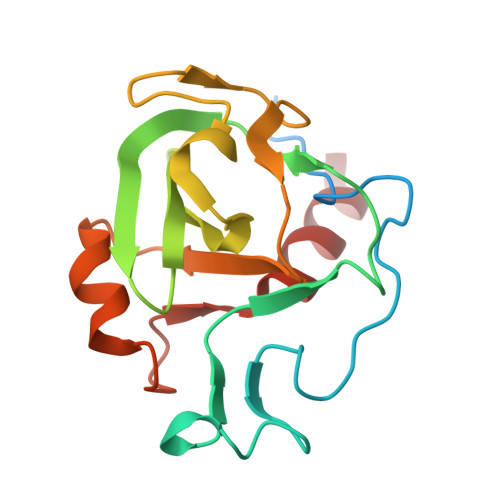
Lactobacillus salivarius REN, a novel probiotic isolated from Chinese centenarians, can adhere to intestinal epithelial cells and subsequently colonize the host. We show here that the surface-layer protein choline-binding protein A (CbpA) of L. salivarius REN was involved in adherence to the human colorectal adenocarcinoma cell line HT-29. Adhesion of a cbpA deletion mutant was significantly reduced compared with that of wild-type, suggesting that CbpA acts as an adhesin that mediates the interaction between the bacterium and its host. To identify the molecular mechanism of adhesion, we determined the crystal structure of a truncated form of CbpA that is likely involved in binding to its cell-surface receptor. The crystal structure identified CbpA as a peptidase of the M23 family whose members harbor a zinc-dependent catalytic site. Therefore, we propose that CbpA acts as a multifunctional surface protein that cleaves the host extracellular matrix and participates in adherence. Moreover, we identified enolase as the CbpA receptor on the surface of HT-29 cells. The present study reveals a new class of surface-layer proteins as well as the molecular mechanism that may contribute to the ability of L. salivarius REN to colonize the human gut.
- Beijing Advanced Innovation Center for Food Nutrition and Human Health, China Agricultural University, Beijing 100083, P. R. China.
Organizational Affiliation: