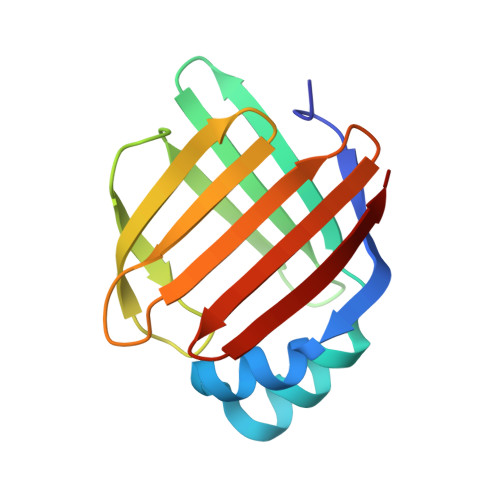
The ligand-mediated affinity of brain-type fatty acid-binding protein for membranes determines the directionality of lipophilic cargo transport.
Cheng, Y.-Y., Huang, Y.-F., Lin, H.-H., Chang, W.W., Lyu, P.-C.(2019) Biochim Biophys Acta Mol Cell Biol Lipids 1864: 158506-158506
- PubMed: 31404652 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbalip.2019.08.002
- Primary Citation Related Structures:
5GGE, 5GKB - PubMed Abstract:
The intracellular transport of lipophilic cargoes is a highly dynamic process. In eukaryotic cells, the uptake and release of long-chain fatty acids (LCFAs) are executed by fatty-acid binding proteins. However, how these carriers control the directionality of cargo trafficking remains unclear. Here, we revealed that the unliganded archetypal Drosophila brain-type fatty acid-binding protein (dFABP) possesses a stronger binding affinity than its liganded counterpart for empty nanodiscs (ND). Titrating unliganded dFABP and nanodiscs with LCFAs rescued the broadening of FABP cross-peak intensities in HSQC spectra from a weakened protein-membrane interaction. Two out of the 3 strongest LCFA contacting residues in dFABP identified by NMR HSQC chemical shift perturbation (CSP) are also part of the 30 ND-contacting residues (out of the total 130 residues in dFABP), revealed by attenuated TROSY signal in the presence of lipid ND to apo-like dFABP. Our crystallographic temperature factor data suggest enhanced αII helix dynamics upon LCFA binding, compensating for the entropic loss in the βC-D/βE-F loops. The aliphatic tail of bound LCFA impedes the charge-charge interaction between dFABP and the head groups of the membrane, and dFABP is prone to dissociate from the membrane upon ligand binding. We therefore conclude that lipophilic ligands participate directly in the control of the functionally required membrane association and dissociation of FABPs.
- Institute of Bioinformatics and Structural Biology, National Tsing Hua University, Hsinchu, Taiwan; National Institute of Cancer Research, National Health Research Institutes, Zhunan, Taiwan.
Organizational Affiliation: