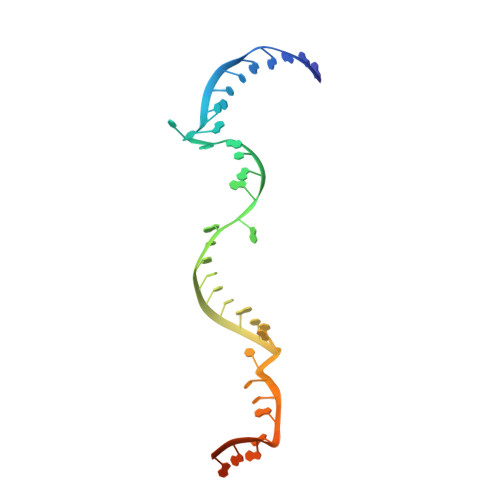
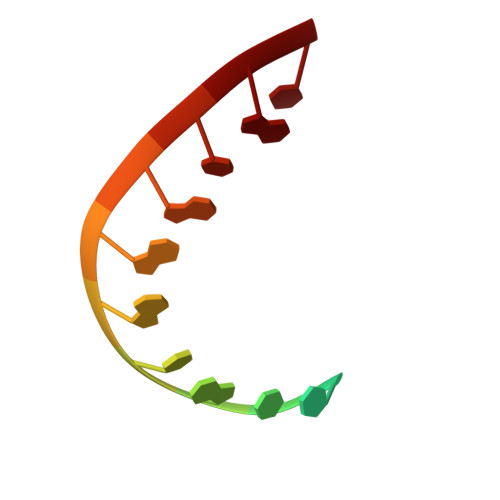
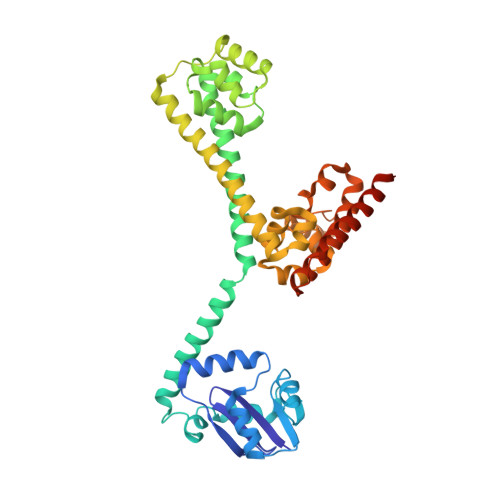
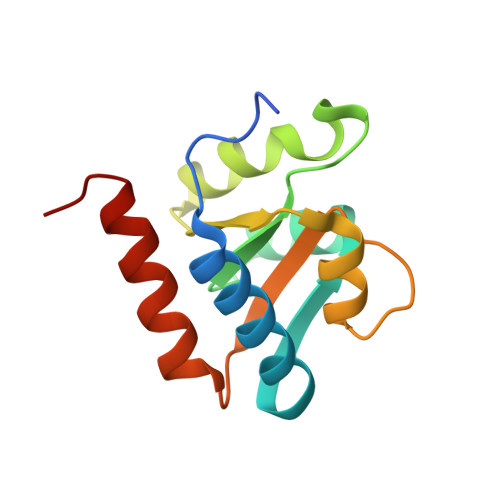
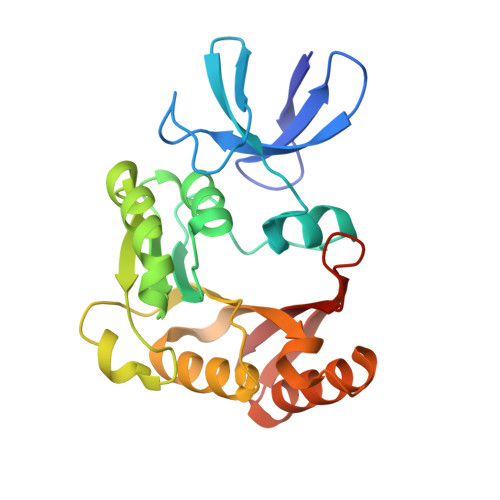
Box C/D guide RNAs recognize a maximum of 10 nt of substrates
Yang, Z., Lin, J., Ye, K.(2016) Proc Natl Acad Sci U S A 113: 10878-10883
- PubMed: 27625427 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1604872113
- Primary Citation Related Structures:
5GIN, 5GIO, 5GIP - PubMed Abstract:
Box C/D RNAs guide site-specific 2'-O-methylation of RNAs in archaea and eukaryotes. The spacer regions between boxes C to D' and boxes C' to D contain the guide sequence that can form a stretch of base pairs with substrate RNAs. The lengths of spacer regions and guide-substrate duplexes are variable among C/D RNAs. In a previously determined structure of C/D ribonucleoprotein (RNP), a 12-nt-long spacer forms 10 bp with the substrate. How spacers and guide-substrate duplexes of other lengths are accommodated remains unknown. Here we analyze how the lengths of spacers and guide-substrate duplexes affect the modification activity and determine three structures of C/D RNPs assembled with different spacers and substrates. We show that the guide can only form a duplex of a maximum of 10 bp with the substrate during modification. Slightly shorter duplexes are tolerated, but longer duplexes must be unwound to fit into a capped protein channel for modification. Spacers with <12 nucleotides are defective, mainly because they cannot load the substrate in the active conformation. For spacers with >12 nucleotides, the excessive unpaired sequences near the box C/C' side are looped out. Our results provide insight into the substrate recognition mechanism of C/D RNA and refute the RNA-swapped model for dimeric C/D RNP.
- College of Biological Sciences, China Agricultural University, Beijing 100193, China; National Institute of Biological Sciences, Beijing 102206, China; Key Laboratory of RNA Biology, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China; Beijing Key Laboratory of Noncoding RNA, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China;
Organizational Affiliation: