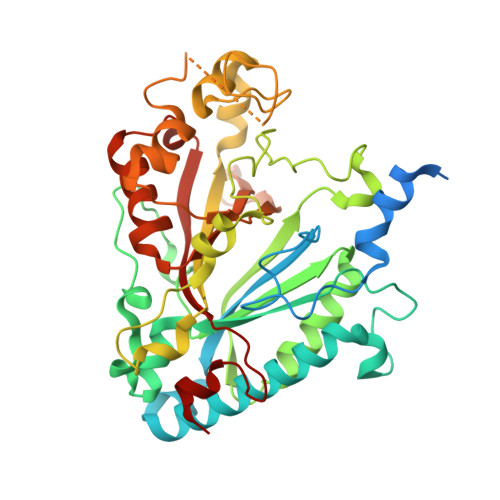
Structure of Thermobifida Fusca Dyp-Type Peroxidase and Activity Towards Kraft Lignin and Lignin Model Compounds.
Rahmanpour, R., Rea, D., Jamshidi, S., Fulop, V., Bugg, T.D.H.(2016) Arch Biochem Biophys 594: 54
- PubMed: 26901432 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2016.02.019
- Primary Citation Related Structures:
5FW4 - PubMed Abstract:
A Dyp-type peroxidase enzyme from thermophilic cellulose degrader Thermobifida fusca (TfuDyP) was investigated for catalytic ability towards lignin oxidation. TfuDyP was characterised kinetically against a range of phenolic substrates, and a compound I reaction intermediate was observed via pre-steady state kinetic analysis at λmax 404 nm. TfuDyP showed reactivity towards Kraft lignin, and was found to oxidise a β-aryl ether lignin model compound, forming an oxidised dimer. A crystal structure of TfuDyP was determined, to 1.8 Å resolution, which was found to contain a diatomic oxygen ligand bound to the heme centre, positioned close to active site residues Asp-203 and Arg-315. The structure contains two channels providing access to the heme cofactor for organic substrates and hydrogen peroxide. Site-directed mutant D203A showed no activity towards phenolic substrates, but reduced activity towards ABTS, while mutant R315Q showed no activity towards phenolic substrates, nor ABTS.
- Department of Chemistry, University of Warwick, Gibbet Hill Road, Coventry CV4 7AL, UK.
Organizational Affiliation: