Crystal Structure of Human Sumo E1 Ufd Domain in Complex with Ubc9.
Liu, B., Castano, L., Lois, M., Reverter, D.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| SUMO-CONJUGATING ENZYME UBC9 | 161 | Homo sapiens | Mutation(s): 0 EC: 6.3.2 (PDB Primary Data), 2.3.2 (UniProt) | 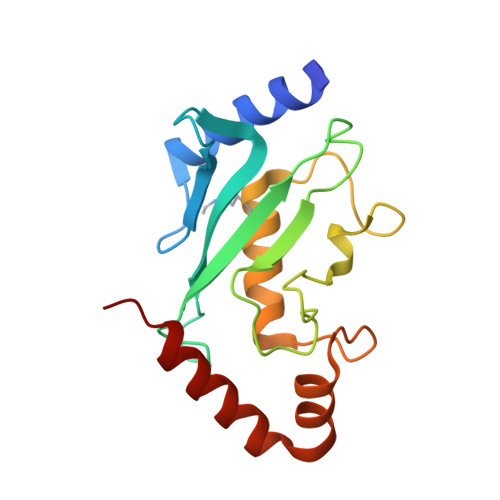 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P63279 GTEx: ENSG00000103275 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63279 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| SUMO-ACTIVATING ENZYME SUBUNIT 2 | 105 | Homo sapiens | Mutation(s): 0 EC: 6.3.2 (PDB Primary Data), 2.3.2 (UniProt) | 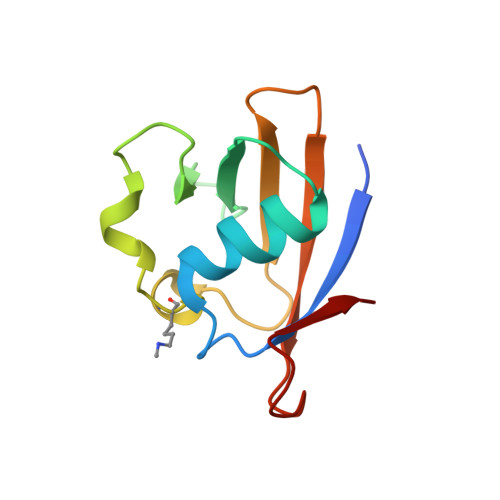 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9UBT2 GTEx: ENSG00000126261 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9UBT2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| LDH Query on LDH | A | L-PEPTIDE LINKING | C8 H18 N2 O2 |  | LYS |
| MLZ Query on MLZ | B | L-PEPTIDE LINKING | C7 H16 N2 O2 |  | LYS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 129.614 | α = 90 |
| b = 129.614 | β = 90 |
| c = 66.578 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| SCALA | data scaling |
| PHASER | phasing |