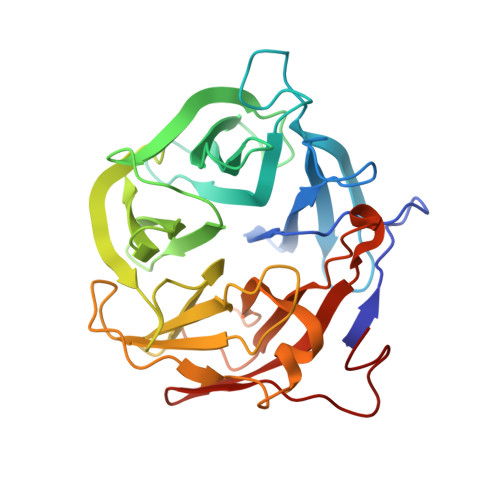
Crystal structure of a putative exo-beta-1,3-galactanase from Bifidobacterium bifidum S17.
Godoy, A.S., de Lima, M.Z., Camilo, C.M., Polikarpov, I.(2016) Acta Crystallogr F Struct Biol Commun 72: 288-293
- PubMed: 27050262 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16003617
- Primary Citation Related Structures:
5FLW - PubMed Abstract:
Given the current interest in second-generation biofuels, carbohydrate-active enzymes have become the most important tool to overcome the structural recalcitrance of the plant cell wall. While some glycoside hydrolase families have been exhaustively described, others remain poorly characterized, especially with regard to structural information. The family 43 glycoside hydrolases are a diverse group of inverting enzymes; the available structure information on these enzymes is mainly from xylosidases and arabinofuranosidase. Currently, only one structure of an exo-β-1,3-galactanase is available. Here, the production, crystallization and structure determination of a putative exo-β-1,3-galactanase from Bifidobacterium bifidum S17 (BbGal43A) are described. BbGal43A was successfully produced and showed activity towards synthetic galactosides. BbGal43A was subsequently crystallized and data were collected to 1.4 Å resolution. The structure shows a single-domain molecule, differing from known homologues, and crystal contact analysis predicts the formation of a dimer in solution. Further biochemical studies are necessary to elucidate the differences between BbGal43A and its characterized homologues.
- Departamento de Física em São Carlos, Universidade de São Paulo, Avenida Trabalhador Saocarlense 400, 13560-970 São Carlos-SP, Brazil.
Organizational Affiliation: