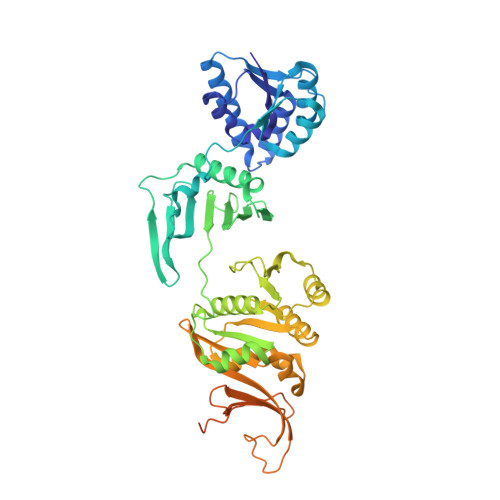
Structure and Function of a Novel ATPase that Interacts with Holliday Junction Resolvase Hjc and Promotes Branch Migration.
Zhai, B., DuPrez, K., Doukov, T.I., Li, H., Huang, M., Shang, G., Ni, J., Gu, L., Shen, Y., Fan, L.(2017) J Mol Biology 429: 1009-1029
- PubMed: 28238763 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2017.02.016
- Primary Citation Related Structures:
5F4H - PubMed Abstract:
Holliday junction (HJ) is a hallmark intermediate in DNA recombination and must be processed by dissolution (for double HJ) or resolution to ensure genome stability. Although HJ resolvases have been identified in all domains of life, there is a long-standing effort to search in prokaryotes and eukarya for proteins promoting HJ migration. Here, we report the structural and functional characterization of a novel ATPase, Sulfolobus islandicusPilT N-terminal-domain-containing ATPase (SisPINA), encoded by the gene adjacent to the resolvase Hjc coding gene. PINA is conserved in archaea and vital for S. islandicus viability. Purified SisPINA forms hexameric rings in the crystalline state and in solution, similar to the HJ migration helicase RuvB in Gram-negative bacteria. Structural analysis suggests that ATP binding and hydrolysis cause conformational changes in SisPINA to drive branch migration. Further studies reveal that SisPINA interacts with SisHjc and coordinates HJ migration and cleavage.
- State Key Laboratory of Microbial Technology, Shandong University, 27 Shanda Nan Road, Jinan 250100, PR China.
Organizational Affiliation: