X-ray structure of the thymidine phosphorylase from Salmonella typhimurium in complex with cytidine and sulphate
Balaev, V.V., Lashkov, A.A., Gabdulkhakov, A.G., Betzel, C., Mikhailov, A.M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Thymidine phosphorylase | 442 | Salmonella enterica subsp. enterica serovar Typhimurium str. LT2 | Mutation(s): 0 Gene Names: deoA, STM4568 EC: 2.4.2.4 | 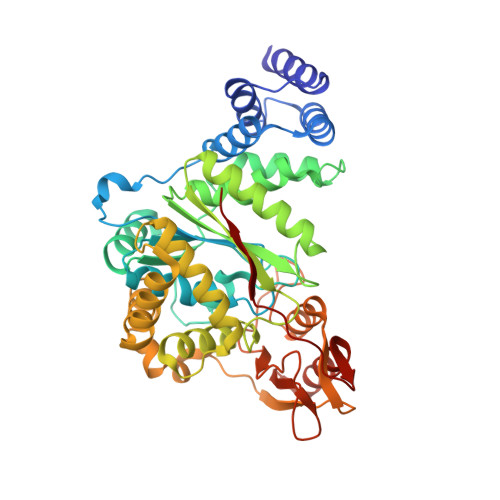 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7CP66 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CTN Download:Ideal Coordinates CCD File | O [auth B] | 4-AMINO-1-BETA-D-RIBOFURANOSYL-2(1H)-PYRIMIDINONE C9 H13 N3 O5 UHDGCWIWMRVCDJ-XVFCMESISA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | C [auth A] D [auth A] E [auth A] F [auth A] G [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | H [auth A], N [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | I [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 193.398 | α = 90 |
| b = 193.398 | β = 90 |
| c = 58.208 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SCALA | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| PHASER | phasing |
| XDS | data reduction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Russian Foundation for Basic Research | Russian Federation | 14-04-00952 |