Optimization of the Expression of Human Aldehyde Oxidase for Investigations of Single-Nucleotide Polymorphisms.
Foti, A., Hartmann, T., Coelho, C., Santos-Silva, T., Romao, M.J., Leimkuhler, S.(2016) Drug Metab Dispos 44: 1277-1285
- PubMed: 26842593 Search on PubMed
- DOI: https://doi.org/10.1124/dmd.115.068395
- Primary Citation Related Structures:
5EPG - PubMed Abstract:
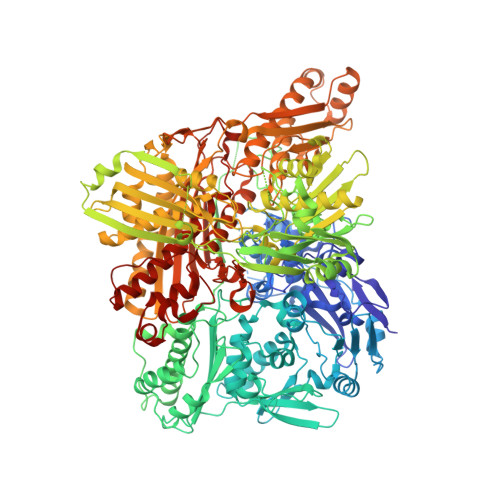
Aldehyde oxidase (AOX1) is an enzyme with broad substrate specificity, catalyzing the oxidation of a wide range of endogenous and exogenous aldehydes as well as N-heterocyclic aromatic compounds. In humans, the enzyme's role in phase I drug metabolism has been established and its importance is now emerging. However, the true physiologic function of AOX1 in mammals is still unknown. Further, numerous single-nucleotide polymorphisms (SNPs) have been identified in human AOX1. SNPs are a major source of interindividual variability in the human population, and SNP-based amino acid exchanges in AOX1 reportedly modulate the catalytic function of the enzyme in either a positive or negative fashion. For the reliable analysis of the effect of amino acid exchanges in human proteins, the existence of reproducible expression systems for the production of active protein in ample amounts for kinetic, spectroscopic, and crystallographic studies is required. In our study we report an optimized expression system for hAOX1 in Escherichia coli using a codon-optimized construct. The codon-optimization resulted in an up to 15-fold increase of protein production and a simplified purification procedure. The optimized expression system was used to study three SNPs that result in amino acid changes C44W, G1269R, and S1271L. In addition, the crystal structure of the S1271L SNP was solved. We demonstrate that the recombinant enzyme can be used for future studies to exploit the role of AOX in drug metabolism, and for the identification and synthesis of new drugs targeting AOX when combined with crystallographic and modeling studies.
- Department of Molecular Enzymology, Institute of Biochemistry and Biology, University of Potsdam, Potsdam, Germany (A.F., T.H., S.L.); UCIBIO-REQUIMTE-Departamento de Química, Faculdade de Ciências e Tecnologia, Universidade NOVA de Lisboa, Caparica, Portugal (C.C., T.S.-S., M.J.R.).
Organizational Affiliation: