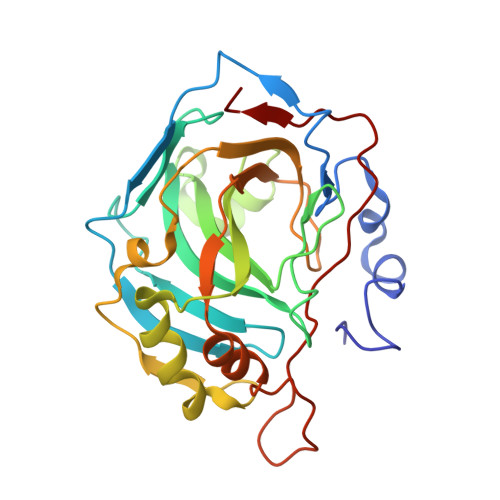
Kinetic and X-ray crystallographic investigations on carbonic anhydrase isoforms I, II, IX and XII of a thioureido analog of SLC-0111.
Lomelino, C.L., Mahon, B.P., McKenna, R., Carta, F., Supuran, C.T.(2016) Bioorg Med Chem 24: 976-981
- PubMed: 26810836 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2016.01.019
- Primary Citation Related Structures:
5EIJ - PubMed Abstract:
SLC-0111 (4-(4-fluorophenylureido)-benzenesulfonamide) is the first carbonic anhydrase (CA, EC 4.2.1.1) IX inhibitor to reach phase I clinical trials as an antitumor/antimetastatic agent. Here we report a kinetic and X-ray crystallographic study of a congener of SLC-0111 which incorporates a thioureido instead of ureido linker between the two aromatic rings as inhibitor of four physiologically relevant CA isoforms. Similar to SLC-0111, the thioureido derivative was a weak hCA I and II inhibitor and a potent one against hCA IX and XII. X-ray crystallography of its adduct with hCA II and comparison of the structure with that of other five hCA II-sulfonamide adducts belonging to the SLC-0111 series, afforded us to understand the particular inhibition profile of the new sulfonamide. Similar to SLC-0111, the thioureido sulfonamide primarily interacted with the hydrophobic side of the hCA II active site, with the tail participating in van der Waals interactions with Phe131 and Pro202, in addition to the coordination of the deprotonated sulfonamide to the active site metal ion. On the contrary, the tail of other sulfonamides belonging to the SLC-0111 series (2-isopropyl-phenyl; 3-nitrophenyl) were orientated towards the hydrophilic half of the active site, which was correlated with orders of magnitude better inhibitory activity against hCA II, and a loss of selectivity for the inhibition of the tumor-associated CAs.
- University of Florida College of Medicine, Department of Biochemistry and Molecular Biology, Gainesville, FL, USA.
Organizational Affiliation: