CSL-RITA complex bound to DNA
Tabaja, N.H., Kovall, R.A.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Recombining binding protein suppressor of hairless | B [auth C] | 422 | Mus musculus | Mutation(s): 0 Gene Names: Rbpj, Igkjrb1, Igkrsbp, Rbpsuh | 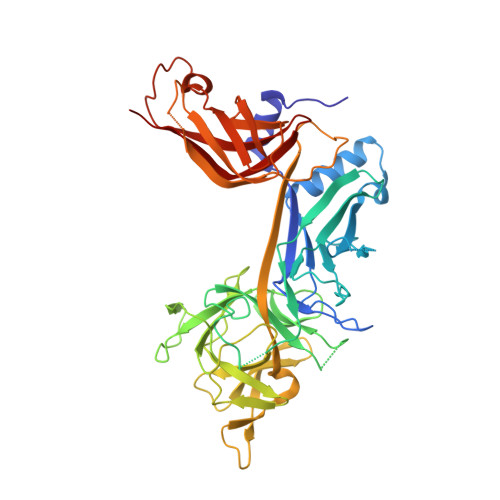 |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:96522 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P31266 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| hRITA | C [auth R] | 16 | synthetic construct | Mutation(s): 0 |  |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q96K30 GTEx: ENSG00000139405 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q96K30 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*AP*AP*TP*CP*TP*TP*TP*CP*CP*CP*AP*CP*AP*GP*T)-3') | A [auth B] | 15 | synthetic construct | 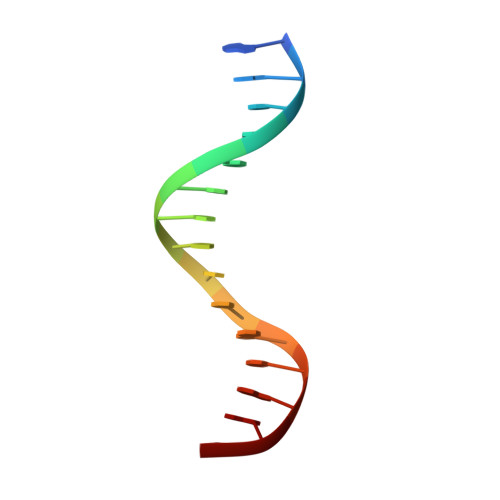 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 4 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*TP*TP*AP*CP*TP*GP*TP*GP*GP*GP*AP*AP*AP*GP*A)-3') | D [auth A] | 15 | synthetic construct |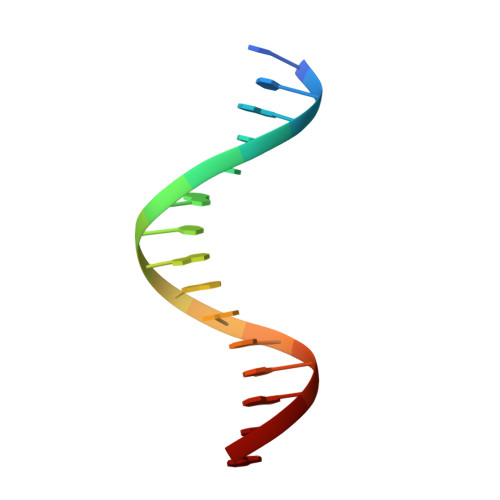  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BU1 Download:Ideal Coordinates CCD File | AA [auth C] BA [auth C] CA [auth C] W [auth C] X [auth C] | 1,4-BUTANEDIOL C4 H10 O2 WERYXYBDKMZEQL-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | DA [auth R] E [auth C] F [auth C] G [auth C] H [auth C] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 76.788 | α = 90 |
| b = 96.41 | β = 90 |
| c = 96.711 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data collection |
| HKL-2000 | data scaling |
| PHASER | phasing |
| PDB_EXTRACT | data extraction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Cancer Institute (NIH/NCI) | United States | R01CA178974-02 |
| National Institutes of Health/National Cancer Institute (NIH/NCI) | United States | T32CA117846-07 |