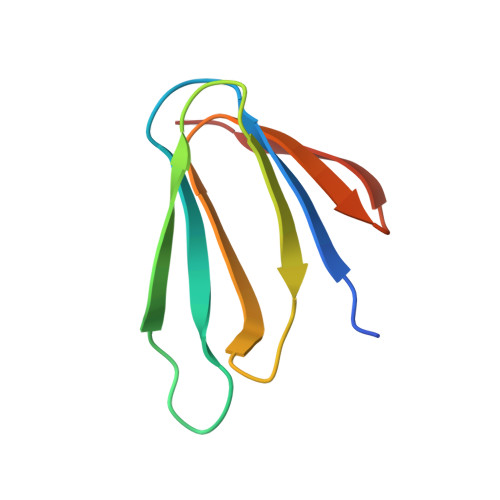
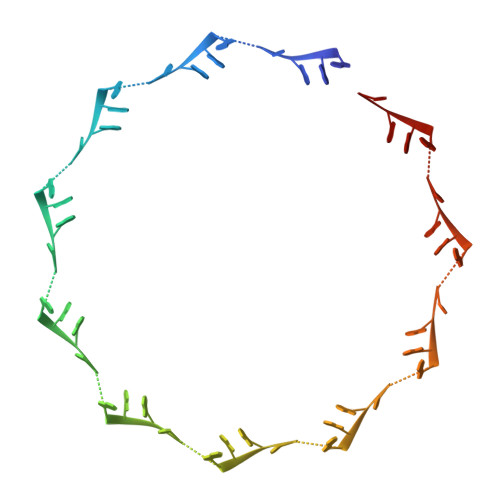
RNA protects a nucleoprotein complex against radiation damage.
Bury, C.S., McGeehan, J.E., Antson, A.A., Carmichael, I., Gerstel, M., Shevtsov, M.B., Garman, E.F.(2016) Acta Crystallogr D Struct Biol 72: 648-657
- PubMed: 27139628 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798316003351
- Primary Citation Related Structures:
5EEU, 5EEV, 5EEW, 5EEX, 5EEY, 5EEZ, 5EF0, 5EF1, 5EF2, 5EF3 - PubMed Abstract:
Radiation damage during macromolecular X-ray crystallographic data collection is still the main impediment for many macromolecular structure determinations. Even when an eventual model results from the crystallographic pipeline, the manifestations of radiation-induced structural and conformation changes, the so-called specific damage, within crystalline macromolecules can lead to false interpretations of biological mechanisms. Although this has been well characterized within protein crystals, far less is known about specific damage effects within the larger class of nucleoprotein complexes. Here, a methodology has been developed whereby per-atom density changes could be quantified with increasing dose over a wide (1.3-25.0 MGy) range and at higher resolution (1.98 Å) than the previous systematic specific damage study on a protein-DNA complex. Specific damage manifestations were determined within the large trp RNA-binding attenuation protein (TRAP) bound to a single-stranded RNA that forms a belt around the protein. Over a large dose range, the RNA was found to be far less susceptible to radiation-induced chemical changes than the protein. The availability of two TRAP molecules in the asymmetric unit, of which only one contained bound RNA, allowed a controlled investigation into the exact role of RNA binding in protein specific damage susceptibility. The 11-fold symmetry within each TRAP ring permitted statistically significant analysis of the Glu and Asp damage patterns, with RNA binding unexpectedly being observed to protect these otherwise highly sensitive residues within the 11 RNA-binding pockets distributed around the outside of the protein molecule. Additionally, the method enabled a quantification of the reduction in radiation-induced Lys and Phe disordering upon RNA binding directly from the electron density.
- Laboratory of Molecular Biophysics, Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, England.
Organizational Affiliation: