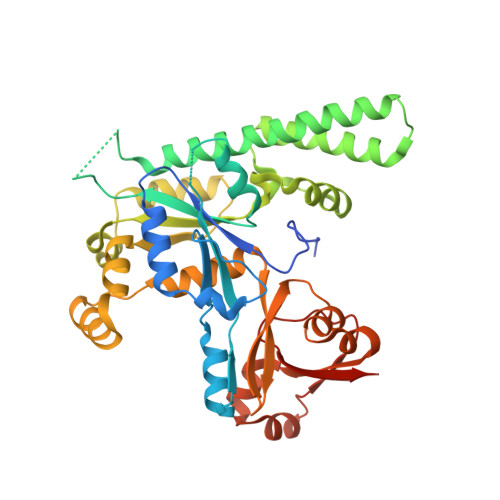
ATP binding by the P-loop NTPase OsYchF1 (an unconventional G protein) contributes to biotic but not abiotic stress responses
Cheung, M.-Y., Li, X., Miao, R., Fong, Y.-H., Li, K.-P., Yung, Y.-L., Yu, M.-H., Wong, K.-B., Chen, Z., Lam, H.-M.(2016) Proc Natl Acad Sci U S A 113: 2648-2653
- PubMed: 26912459 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1522966113
- Primary Citation Related Structures:
5EE0, 5EE1, 5EE3, 5EE9 - PubMed Abstract:
G proteins are involved in almost all aspects of the cellular regulatory pathways through their ability to bind and hydrolyze GTP. The YchF subfamily, interestingly, possesses the unique ability to bind both ATP and GTP, and is possibly an ancestral form of G proteins based on phylogenetic studies and is present in all kingdoms of life. However, the biological significance of such a relaxed ligand specificity has long eluded researchers. Here, we have elucidated the different conformational changes caused by the binding of a YchF homolog in rice (OsYchF1) to ATP versus GTP by X-ray crystallography. Furthermore, by comparing the 3D relationships of the ligand position and the various amino acid residues at the binding sites in the crystal structures of the apo-bound and ligand-bound versions, a mechanism for the protein's ability to bind both ligands is revealed. Mutation of the noncanonical G4 motif of the OsYchF1 to the canonical sequence for GTP specificity precludes the binding/hydrolysis of ATP and prevents OsYchF1 from functioning as a negative regulator of plant-defense responses, while retaining its ability to bind/hydrolyze GTP and its function as a negative regulator of abiotic stress responses, demonstrating the specific role of ATP-binding/hydrolysis in disease resistance. This discovery will have a significant impact on our understanding of the structure-function relationships of the YchF subfamily of G proteins in all kingdoms of life.
- School of Life Sciences, The Chinese University of Hong Kong, Shatin, N.T., Hong Kong SAR; Center for Soybean Research of the State Key Laboratory of Agrobiotechnology, The Chinese University of Hong Kong, Shatin, N.T., Hong Kong SAR;
Organizational Affiliation: