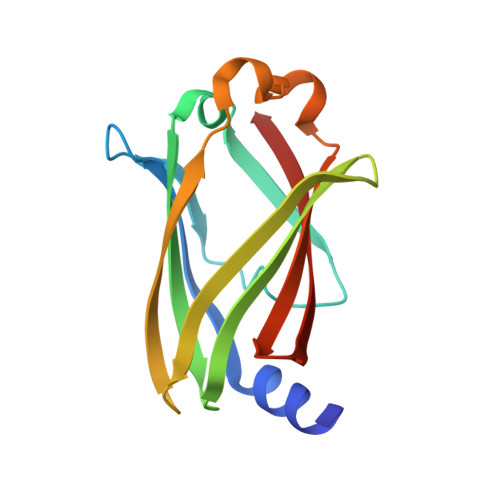
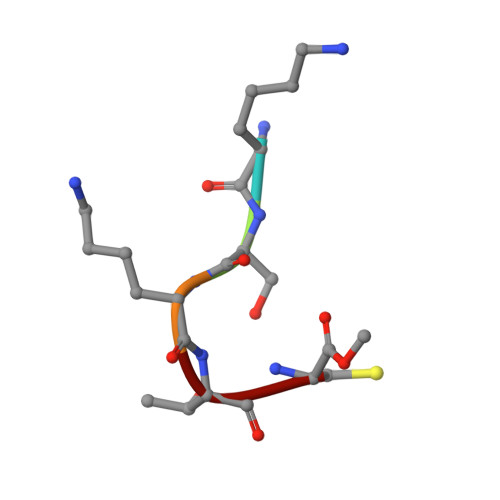
The N- and C-terminal ends of RPGR can bind to PDE6 delta.
Fansa, E.K., O'Reilly, N.J., Ismail, S., Wittinghofer, A.(2015) EMBO Rep 16: 1583-1585
- PubMed: 26553937 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embr.201541404
- Primary Citation Related Structures:
5E8F - PubMed Abstract:
This study shows that the prenylated C‐terminus of RPGR can bind to PDE6δ with high affinity, suggesting two distinct binding sites of the RPGR/PDE6δ complex. The serine residue at the −3 position relative to the prenylated cysteine seems to play a key role in defining the selectivity of PDE6δ towards ciliary prenylated cargo. [Image: see text]
- Max Planck Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: