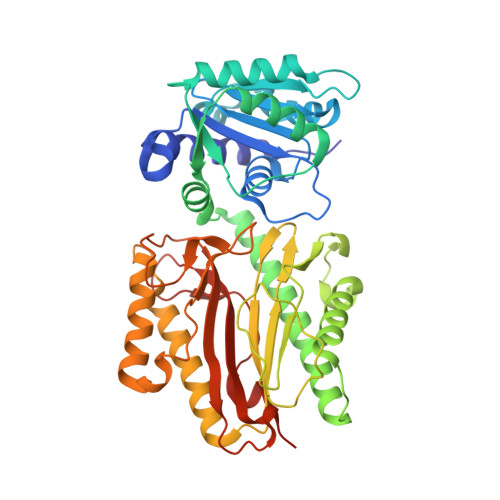
Structure of the human histone chaperone FACT Spt16 N-terminal domain.
Marciano, G., Huang, D.T.(2016) Acta Crystallogr F Struct Biol Commun 72: 121-128
- PubMed: 26841762 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15024565
- Primary Citation Related Structures:
5E5B - PubMed Abstract:
The histone chaperone FACT plays an important role in facilitating nucleosome assembly and disassembly during transcription. FACT is a heterodimeric complex consisting of Spt16 and SSRP1. The N-terminal domain of Spt16 resembles an inactive aminopeptidase. How this domain contributes to the histone chaperone activity of FACT remains elusive. Here, the crystal structure of the N-terminal domain (NTD) of human Spt16 is reported at a resolution of 1.84 Å. The structure adopts an aminopeptidase-like fold similar to those of the Saccharomyces cerevisiae and Schizosaccharomyces pombe Spt16 NTDs. Isothermal titration calorimetry analyses show that human Spt16 NTD binds histones H3/H4 with low-micromolar affinity, suggesting that Spt16 NTD may contribute to histone binding in the FACT complex. Surface-residue conservation and electrostatic analysis reveal a conserved acidic patch that may be involved in histone binding.
- Cancer Research UK Beatson Institute, Garscube Estate, Switchback Road, Glasgow G61 1BD, Scotland.
Organizational Affiliation: