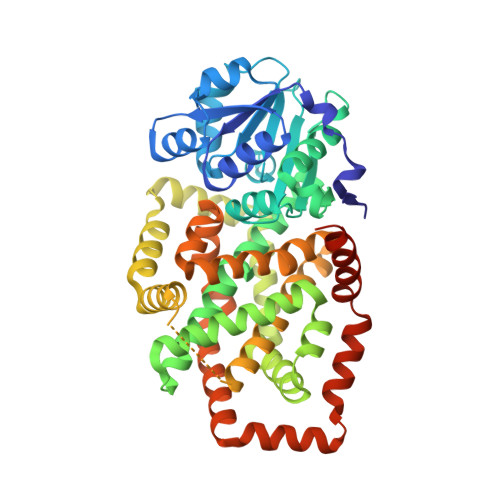
Artificial domain duplication replicates evolutionary history of ketol-acid reductoisomerases.
Cahn, J.K., Brinkmann-Chen, S., Buller, A.R., Arnold, F.H.(2016) Protein Sci 25: 1241-1248
- PubMed: 26644020 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2852
- Primary Citation Related Structures:
5E4R - PubMed Abstract:
The duplication of protein structural domains has been proposed as a common mechanism for the generation of new protein folds. A particularly interesting case is the class II ketol-acid reductoisomerase (KARI), which putatively arose from an ancestral class I KARI by duplication of the C-terminal domain and corresponding loss of obligate dimerization. As a result, the class II enzymes acquired a deeply embedded figure-of-eight knot. To test this evolutionary hypothesis we constructed a novel class II KARI by duplicating the C-terminal domain of a hyperthermostable class I KARI. The new protein is monomeric, as confirmed by gel filtration and X-ray crystallography, and has the deeply knotted class II KARI fold. Surprisingly, its catalytic activity is nearly unchanged from the parent KARI. This provides strong evidence in support of domain duplication as the mechanism for the evolution of the class II KARI fold and demonstrates the ability of domain duplication to generate topological novelty in a function-neutral manner.
- Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, California, 91125.
Organizational Affiliation: