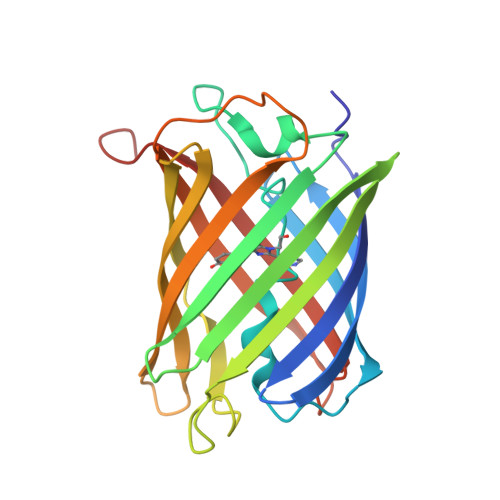
Arginine 66 Controls Dark-State Formation in Green-to-Red Photoconvertible Fluorescent Proteins.
Berardozzi, R., Adam, V., Martins, A., Bourgeois, D.(2016) J Am Chem Soc 138: 558-565
- PubMed: 26675944 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.5b09923
- Primary Citation Related Structures:
5DTL - PubMed Abstract:
Photoactivated localization microscopy (PALM) is a powerful technique to investigate cellular nanostructures quantitatively and dynamically. However, the use of PALM for molecular counting or single-particle tracking remains limited by the propensity of photoconvertible fluorescent protein markers (PCFPs) to repeatedly enter dark states. By designing the single mutants mEos2-A69T and Dendra2-T69A, we completely swapped the blinking behaviors of mEos2 and Dendra2, two popular PCFPs. We combined X-ray crystallography and single-molecule microscopy to show that blinking in mEos2 and Dendra2 is largely controlled by the orientation of arginine 66, a highly conserved residue in Anthozoan PCFPs. The Arg66 side-chain conformation affects the bleaching and the on-to-off transition quantum yields, as well as the fraction of molecules entering long-lived dark states, resulting in widely different apparent blinking behaviors that largely modulate the efficiency of current blinking correction procedures. The present work provides mechanistic insight into the complex photophysics of Anthozoan PCFPs and will facilitate future engineering of bright and low-blinking variants suitable for PALM.
- Institut de Biologie Structurale, Université Grenoble Alpes , CEA, CNRS, 38044 Grenoble, France.
Organizational Affiliation: