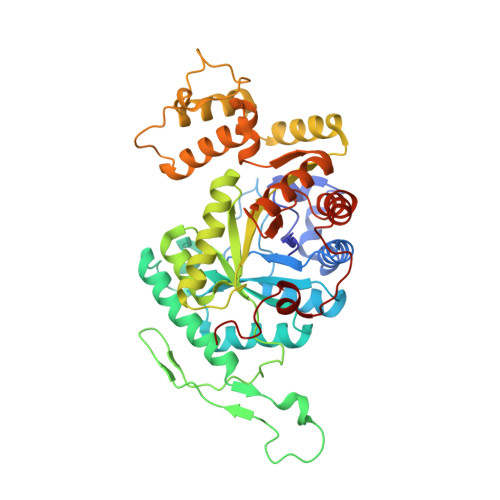
Structural and biochemical characterization of EDTA monooxygenase and its physical interaction with a partner flavin reductase.
Jun, S.Y., Lewis, K.M., Youn, B., Xun, L., Kang, C.(2016) Mol Microbiol 100: 989-1003
- PubMed: 26928990 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/mmi.13363
- Primary Citation Related Structures:
5DQP - PubMed Abstract:
Ethylenediaminetetraacetate (EDTA) is currently the most abundant organic pollutant due to its recalcitrance and extensive use. Only a few bacteria can degrade it, using EDTA monooxygenase (EmoA) to initiate the degradation. EmoA is an FMNH2 -dependent monooxygenase that requires an NADH:FMN oxidoreductase (EmoB) to provide FMNH2 as a cosubstrate. Although EmoA has been identified from Chelativorans (ex. Mesorhizobium) sp. BNC1, its catalytic mechanism is unknown. Crystal structures of EmoA revealed a domain-like insertion into a TIM-barrel, which might serve as a flexible lid for the active site. Docking of MgEDTA(2-) into EmoA identified an intricate hydrogen bond network connected to Tyr(71) , which should potentially lower its pKa. Tyr(71) , along with nearby Glu(70) and a peroxy flavin, facilitates a keto-enol transition of the leaving acetyl group of EDTA. Further, for the first time, the physical interaction between EmoA and EmoB was observed by ITC, molecular docking and enzyme kinetic assay, which enhanced both EmoA and EmoB activities probably through coupled channelling of FMNH2 .
- Department of Chemistry, Washington State University, Pullman, WA, 99164-4630, USA.
Organizational Affiliation: