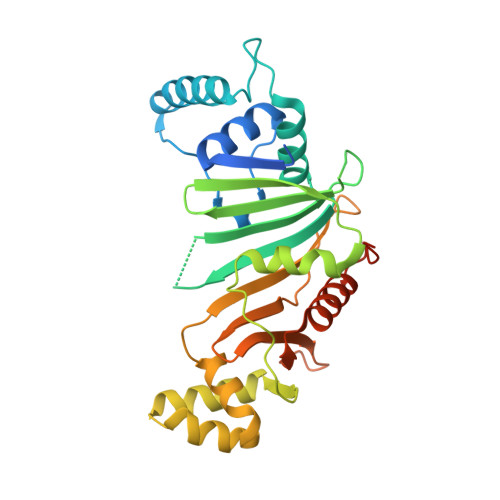
Structural Variability of EspG Chaperones from Mycobacterial ESX-1, ESX-3, and ESX-5 Type VII Secretion Systems.
Tuukkanen, A.T., Freire, D., Chan, S., Arbing, M.A., Reed, R.W., Evans, T.J., Zenkeviciute, G., Kim, J., Kahng, S., Sawaya, M.R., Chaton, C.T., Wilmanns, M., Eisenberg, D., Parret, A.H.A., Korotkov, K.V.(2019) J Mol Biology 431: 289-307
- PubMed: 30419243 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2018.11.003
- Primary Citation Related Structures:
4L4W, 4RCL, 5DLB, 5SXL, 5VBA - PubMed Abstract:
Type VII secretion systems (ESX) are responsible for transport of multiple proteins in mycobacteria. How different ESX systems achieve specific secretion of cognate substrates remains elusive. In the ESX systems, the cytoplasmic chaperone EspG forms complexes with heterodimeric PE-PPE substrates that are secreted from the cells or remain associated with the cell surface. Here we report the crystal structure of the EspG 1 chaperone from the ESX-1 system determined using a fusion strategy with T4 lysozyme. EspG 1 adopts a quasi 2-fold symmetric structure that consists of a central β-sheet and two α-helical bundles. In addition, we describe the structures of EspG 3 chaperones from four different crystal forms. Alternate conformations of the putative PE-PPE binding site are revealed by comparison of the available EspG 3 structures. Analysis of EspG 1 , EspG 3 , and EspG 5 chaperones using small-angle X-ray scattering reveals that EspG 1 and EspG 3 chaperones form dimers in solution, which we observed in several of our crystal forms. Finally, we propose a model of the ESX-3 specific EspG 3 -PE5-PPE4 complex based on the small-angle X-ray scattering analysis.
- European Molecular Biology Laboratory, Hamburg Unit, Hamburg 22607, Germany.
Organizational Affiliation: