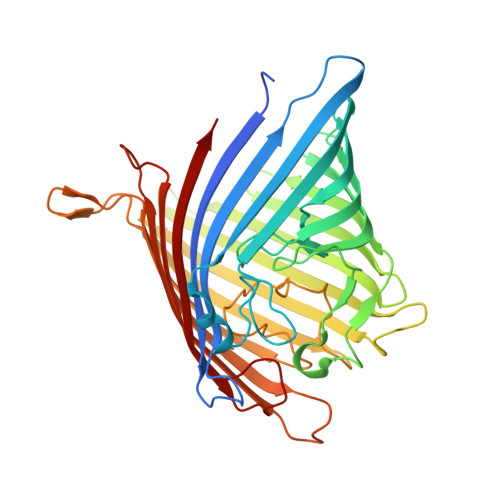
Structural Insights into Outer Membrane Permeability of Acinetobacter baumannii.
Zahn, M., Bhamidimarri, S.P., Basle, A., Winterhalter, M., van den Berg, B.(2016) Structure 24: 221-231
- PubMed: 26805524 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2015.12.009
- Primary Citation Related Structures:
5DL5, 5DL6, 5DL7, 5DL8 - PubMed Abstract:
Bacterial resistance against antibiotics is an increasing global health problem. In Gram-negative bacteria the low permeability of the outer membrane (OM) is a major factor contributing to resistance, making it important to understand channel-mediated small-molecule passage of the OM. Acinetobacter baumannii has five Occ (OM carboxylate channel) proteins, which collectively are of major importance for the entry of small molecules. To improve our understanding of the OM permeability of A. baumannii, we present here the X-ray crystal structures of four Occ proteins, renamed OccAB1 to OccAB4. In addition we have carried out a biochemical and biophysical characterization using electrophysiology and liposome swelling experiments, providing information on substrate specificities. We identify OccAB1 as having the largest pore of the Occ proteins with corresponding high rates of small-molecule uptake, and we suggest that the future design of efficient antibiotics should focus on scaffolds that can permeate efficiently through the OccAB1 channel.
- Institute for Cellular and Molecular Biosciences, Medical School, Newcastle University, Newcastle upon Tyne NE2 4HH, UK.
Organizational Affiliation: