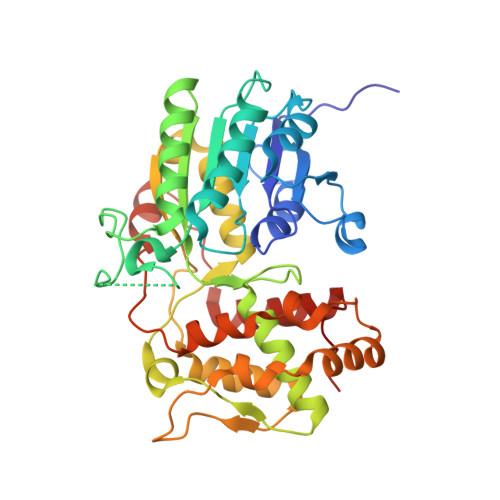
Structural determinants of reductive terpene cyclization in iridoid biosynthesis.
Kries, H., Caputi, L., Stevenson, C.E., Kamileen, M.O., Sherden, N.H., Geu-Flores, F., Lawson, D.M., O'Connor, S.E.(2016) Nat Chem Biol 12: 6-8
- PubMed: 26551396 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1955
- Primary Citation Related Structures:
5DCU, 5DCW, 5DCY, 5DF1 - PubMed Abstract:
The carbon skeleton of ecologically and pharmacologically important iridoid monoterpenes is formed in a reductive cyclization reaction unrelated to canonical terpene cyclization. Here we report the crystal structure of the recently discovered iridoid cyclase (from Catharanthus roseus) bound to a mechanism-inspired inhibitor that illuminates substrate binding and catalytic function of the enzyme. Key features that distinguish iridoid synthase from its close homolog progesterone 5β-reductase are highlighted.
- The John Innes Centre, Department of Biological Chemistry, Norwich Research Park, Norwich NR4 7UH, UK.
Organizational Affiliation: