CRYSTAL STRUCTURE OF THE BASE OF THE RIBOSOMAL P STALK FROM METHANOCOCCUS JANNASCHII WITH ANTIBIOTIC THIOSTREPTON
Gabdulkhakov, A.G., Mitroshin, I.V., Garber, M.B.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 50S ribosomal protein L10 | 213 | Methanocaldococcus jannaschii | Mutation(s): 1 Gene Names: rpl10, rplP0, MJ0509 | 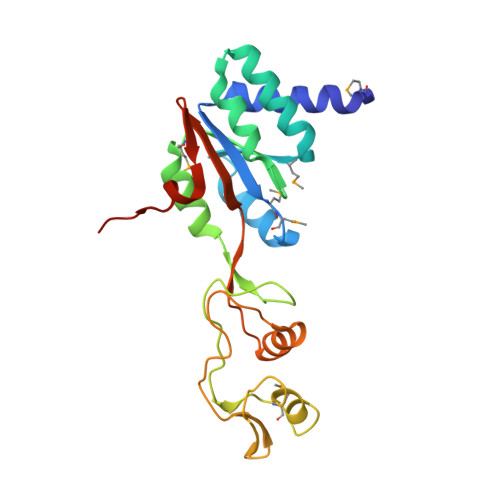 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P54049 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 50S ribosomal protein L11 | 161 | Methanocaldococcus jannaschii | Mutation(s): 0 Gene Names: rpl11, MJ0373 | 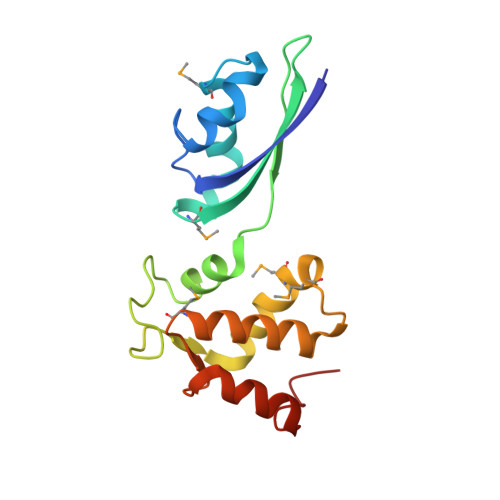 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P54030 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| THIOSTREPTON | 19 | Streptomyces azureus | Mutation(s): 0 | 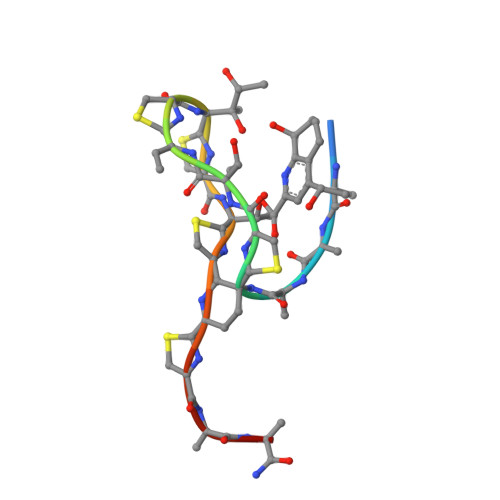 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0C8P8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| 23S ribosomal RNA | 74 | Methanocaldococcus jannaschii | 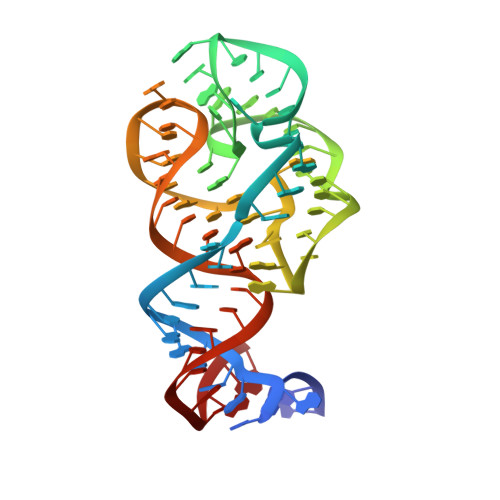 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| TRS Download:Ideal Coordinates CCD File | R [auth A], S [auth A], U [auth B] | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL C4 H12 N O3 LENZDBCJOHFCAS-UHFFFAOYSA-O |  | ||
| MG Download:Ideal Coordinates CCD File | E [auth A] F [auth A] G [auth A] H [auth A] I [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| NA Download:Ideal Coordinates CCD File | O [auth A], P [auth A], Q [auth A], T [auth B] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Modified Residues 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | B | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| BB9 Query on BB9 | D | PEPTIDE LINKING | C3 H5 N O2 S |  | CYS |
| DBU Query on DBU | D | PEPTIDE LINKING | C4 H7 N O2 |  | THR |
| DCY Query on DCY | D | D-PEPTIDE LINKING | C3 H7 N O2 S |  | -- |
| DHA Query on DHA | D | PEPTIDE LINKING | C3 H5 N O2 |  | SER |
| MH6 Query on MH6 | D | PEPTIDE LINKING | C3 H5 N O3 |  | SER |
| TS9 Query on TS9 | D | L-PEPTIDE LINKING | C6 H13 N O4 |  | ILE |
| Entity ID: 4 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_000223 Query on PRD_000223 | D | THIOSTREPTON | Thiopeptide / Antibiotic |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 125.07 | α = 90 |
| b = 127.16 | β = 90 |
| c = 132.93 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| PHENIX | phasing |