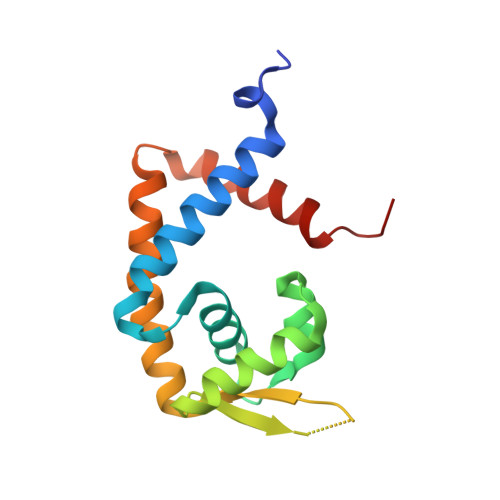
The activity of CouR, a MarR family transcriptional regulator, is modulated through a novel molecular mechanism.
Otani, H., Stogios, P.J., Xu, X., Nocek, B., Li, S.N., Savchenko, A., Eltis, L.D.(2016) Nucleic Acids Res 44: 595-607
- PubMed: 26400178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkv955
- Primary Citation Related Structures:
5CYV - PubMed Abstract:
CouR, a MarR-type transcriptional repressor, regulates the cou genes, encoding p-hydroxycinnamate catabolism in the soil bacterium Rhodococcus jostii RHA1. The CouR dimer bound two molecules of the catabolite p-coumaroyl-CoA (Kd = 11 ± 1 μM). The presence of p-coumaroyl-CoA, but neither p-coumarate nor CoASH, abrogated CouR's binding to its operator DNA in vitro. The crystal structures of ligand-free CouR and its p-coumaroyl-CoA-bound form showed no significant conformational differences, in contrast to other MarR regulators. The CouR-p-coumaroyl-CoA structure revealed two ligand molecules bound to the CouR dimer with their phenolic moieties occupying equivalent hydrophobic pockets in each protomer and their CoA moieties adopting non-equivalent positions to mask the regulator's predicted DNA-binding surface. More specifically, the CoA phosphates formed salt bridges with predicted DNA-binding residues Arg36 and Arg38, changing the overall charge of the DNA-binding surface. The substitution of either arginine with alanine completely abrogated the ability of CouR to bind DNA. By contrast, the R36A/R38A double variant retained a relatively high affinity for p-coumaroyl-CoA (Kd = 89 ± 6 μM). Together, our data point to a novel mechanism of action in which the ligand abrogates the repressor's ability to bind DNA by steric occlusion of key DNA-binding residues and charge repulsion of the DNA backbone.
- Department of Microbiology and Immunology, Life Sciences Institute, The University of British Columbia, Vancouver, British Columbia V6T 1Z3, Canada.
Organizational Affiliation: