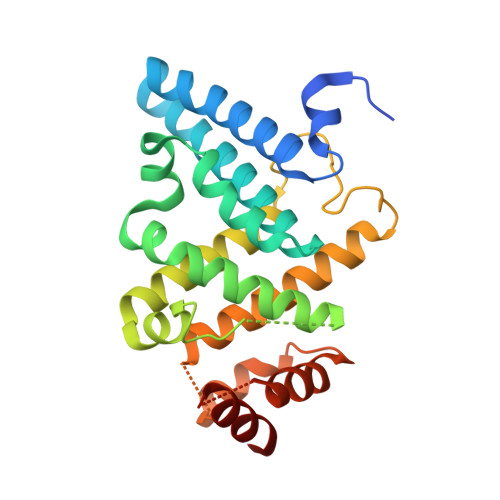
PGL germ granule assembly protein is a base-specific, single-stranded RNase.
Aoki, S.T., Kershner, A.M., Bingman, C.A., Wickens, M., Kimble, J.(2016) Proc Natl Acad Sci U S A 113: 1279-1284
- PubMed: 26787882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1524400113
- Primary Citation Related Structures:
5COW, 5CV1, 5CV3 - PubMed Abstract:
Cellular RNA-protein (RNP) granules are ubiquitous and have fundamental roles in biology and RNA metabolism, but the molecular basis of their structure, assembly, and function is poorly understood. Using nematode "P-granules" as a paradigm, we focus on the PGL granule scaffold protein to gain molecular insights into RNP granule structure and assembly. We first identify a PGL dimerization domain (DD) and determine its crystal structure. PGL-1 DD has a novel 13 α-helix fold that creates a positively charged channel as a homodimer. We investigate its capacity to bind RNA and discover unexpectedly that PGL-1 DD is a guanosine-specific, single-stranded endonuclease. Discovery of the PGL homodimer, together with previous results, suggests a model in which the PGL DD dimer forms a fundamental building block for P-granule assembly. Discovery of the PGL RNase activity expands the role of RNP granule assembly proteins to include enzymatic activity in addition to their job as structural scaffolds.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, WI 53706;
Organizational Affiliation: