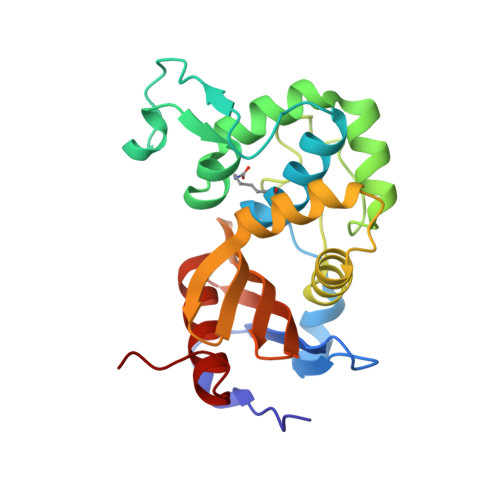
Class D beta-lactamases do exist in Gram-positive bacteria.
Toth, M., Antunes, N.T., Stewart, N.K., Frase, H., Bhattacharya, M., Smith, C.A., Vakulenko, S.B.(2016) Nat Chem Biol 12: 9-14
- PubMed: 26551395 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1950
- Primary Citation Related Structures:
5CTM, 5CTN - PubMed Abstract:
Production of β-lactamases of one of four molecular classes (A, B, C and D) is the major mechanism of bacterial resistance to β-lactams, the largest class of antibiotics, which have saved countless lives since their inception 70 years ago. Although several hundred efficient class D enzymes have been identified in Gram-negative pathogens over the last four decades, none have been reported in Gram-positive bacteria. Here we demonstrate that efficient class D β-lactamases capable of hydrolyzing a wide array of β-lactam substrates are widely disseminated in various species of environmental Gram-positive organisms. Class D enzymes of Gram-positive bacteria have a distinct structural architecture and employ a unique substrate-binding mode that is quite different from that of all currently known class A, C and D β-lactamases. These enzymes thus constitute a previously unknown reservoir of novel antibiotic-resistance enzymes.
- Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, Indiana, USA.
Organizational Affiliation: