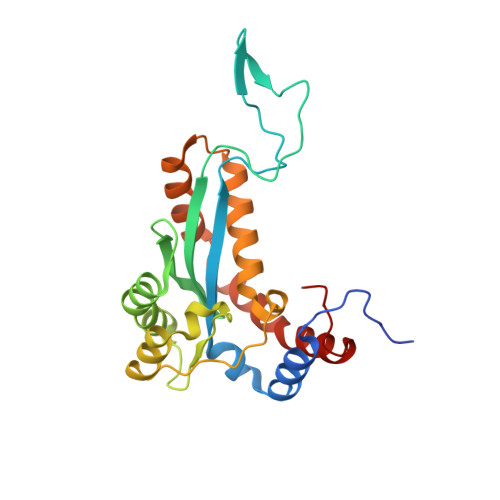
Sleeping Beauty transposase structure allows rational design of hyperactive variants for genetic engineering.
Voigt, F., Wiedemann, L., Zuliani, C., Querques, I., Sebe, A., Mates, L., Izsvak, Z., Ivics, Z., Barabas, O.(2016) Nat Commun 7: 11126-11126
- PubMed: 27025571 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11126
- Primary Citation Related Structures:
5CR4 - PubMed Abstract:
Sleeping Beauty (SB) is a prominent Tc1/mariner superfamily DNA transposon that provides a popular genome engineering tool in a broad range of organisms. It is mobilized by a transposase enzyme that catalyses DNA cleavage and integration at short specific sequences at the transposon ends. To facilitate SB's applications, here we determine the crystal structure of the transposase catalytic domain and use it to model the SB transposase/transposon end/target DNA complex. Together with biochemical and cell-based transposition assays, our structure reveals mechanistic insights into SB transposition and rationalizes previous hyperactive transposase mutations. Moreover, our data enables us to design two additional hyperactive transposase variants. Our work provides a useful resource and proof-of-concept for structure-based engineering of tailored SB transposases.
- European Molecular Biology Laboratory, Structural and Computational Biology Unit, Meyerhofstrasse 1, Heidelberg 69117, Germany.
Organizational Affiliation: