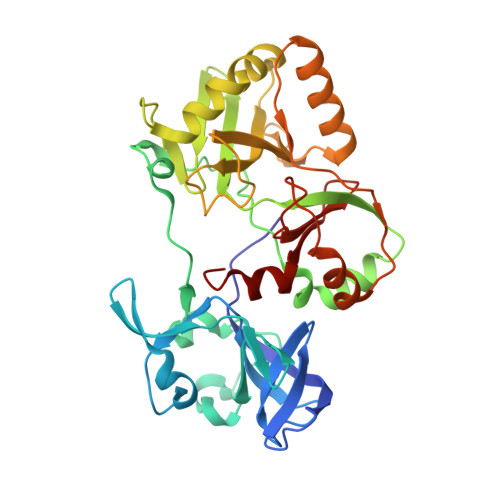
Structure and two-metal mechanism of a eukaryal nick-sealing RNA ligase.
Unciuleac, M.C., Goldgur, Y., Shuman, S.(2015) Proc Natl Acad Sci U S A 112: 13868-13873
- PubMed: 26512110 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1516536112
- Primary Citation Related Structures:
5COT, 5COU, 5COV - PubMed Abstract:
ATP-dependent RNA ligases are agents of RNA repair that join 3'-OH and 5'-PO4 RNA ends. Naegleria gruberi RNA ligase (NgrRnl) exemplifies a family of RNA nick-sealing enzymes found in bacteria, viruses, and eukarya. Crystal structures of NgrRnl at three discrete steps along the reaction pathway-covalent ligase-(lysyl-Nζ)-AMP•Mn(2+) intermediate; ligase•ATP•(Mn(2+))2 Michaelis complex; and ligase•Mn(2+) complex-highlight a two-metal mechanism of nucleotidyl transfer, whereby (i) an enzyme-bound "catalytic" metal coordination complex lowers the pKa of the lysine nucleophile and stabilizes the transition state of the ATP α phosphate; and (ii) a second metal coordination complex bridges the β- and γ-phosphates. The NgrRnl N domain is a distinctively embellished oligonucleotide-binding (OB) fold that engages the γ-phosphate and associated metal complex and orients the pyrophosphate leaving group for in-line catalysis with stereochemical inversion at the AMP phosphate. The unique domain architecture of NgrRnl fortifies the theme that RNA ligases have evolved many times, and independently, by fusions of a shared nucleotidyltransferase domain to structurally diverse flanking modules. The mechanistic insights to lysine adenylylation gained from the NgrRnl structures are likely to apply broadly to the covalent nucleotidyltransferase superfamily of RNA ligases, DNA ligases, and RNA capping enzymes.
- Molecular Biology Program, Sloan-Kettering Institute, New York, NY 10065;
Organizational Affiliation: