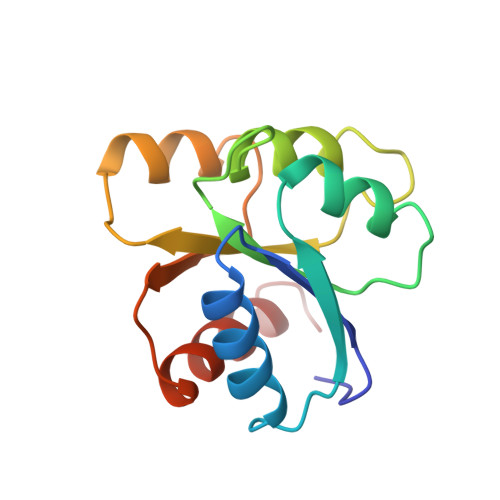
Crystal structures of CheY mutants Y106W and T87I/Y106W. CheY activation correlates with movement of residue 106.
Zhu, X., Rebello, J., Matsumura, P., Volz, K.(1997) J Biological Chem 272: 5000-5006
- PubMed: 9030562 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.272.8.5000
- Primary Citation Related Structures:
5CHY, 6CHY - PubMed Abstract:
Position 106 in CheY is highly conserved as an aromatic residue in the response regulator superfamily. In the structure of the wild-type, apo-CheY, Tyr106 is a rotamer whose electron density is observed in both the inside and the outside positions. In the structure of the T87I mutant of CheY, the threonine to isoleucine change at position 87 causes the side chain of Tyr106 to be exclusively restricted to the outside position. In this report we demonstrate that the T87I mutation causes cells to be smooth swimming and non-chemotactic. We also show that another CheY mutant, Y106W, causes cells to be more tumbly than wild-type CheY, and impairs chemotaxis. In the structure of Y106W, the side chain of Trp106 stays exclusively in the inside position. Furthermore, a T87I/Y106W double mutant, which confers the same phenotype as T87I, restricts the side chain of Trp106 to the outside position. The results from these behavioral and structural studies indicate that the rotameric nature of the Tyr106 residue is involved in activation of the CheY molecule. Specifically, CheY's signaling ability correlates with the conformational heterogeneity of the Tyr106 side chain. Our data also suggest that these mutations affect the signal at an event subsequent to phosphorylation.
- Department of Microbiology and Immunology, University of Illinois, Chicago, Illinois 60612, USA.
Organizational Affiliation: