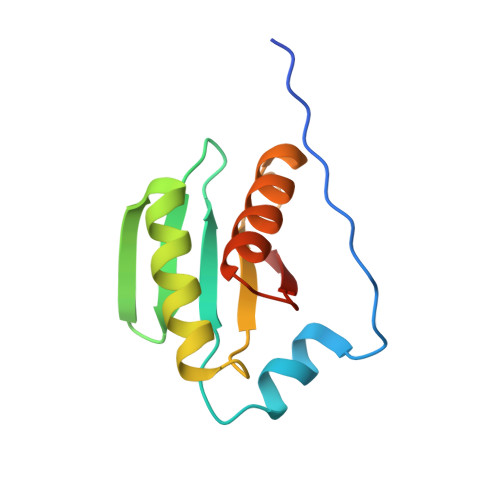
Structure and Function of the Escherichia coli Tol-Pal Stator Protein TolR.
Wojdyla, J.A., Cutts, E., Kaminska, R., Papadakos, G., Hopper, J.T., Stansfeld, P.J., Staunton, D., Robinson, C.V., Kleanthous, C.(2015) J Biological Chem 290: 26675-26687
- PubMed: 26354441 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.671586
- Primary Citation Related Structures:
5BY4 - PubMed Abstract:
TolR is a 15-kDa inner membrane protein subunit of the Tol-Pal complex in Gram-negative bacteria, and its function is poorly understood. Tol-Pal is recruited to cell division sites where it is involved in maintaining the integrity of the outer membrane. TolR is related to MotB, the peptidoglycan (PG)-binding stator protein from the flagellum, suggesting it might serve a similar role in Tol-Pal. The only structure thus far reported for TolR is of the periplasmic domain from Haemophilus influenzae in which N- and C-terminal residues had been deleted (TolR(62-133), Escherichia coli numbering). H. influenzae TolR(62-133) is a symmetrical dimer with a large deep cleft at the dimer interface. Here, we present the 1.7-Å crystal structure of the intact periplasmic domain of E. coli TolR (TolR(36-142)). E. coli TolR(36-142) is also dimeric, but the architecture of the dimer is radically different from that of TolR(62-133) due to the intertwining of its N and C termini. TolR monomers are rotated ∼180° relative to each other as a result of this strand swapping, obliterating the putative PG-binding groove seen in TolR(62-133). We found that removal of the strand-swapped regions (TolR(60-133)) exposes cryptic PG binding activity that is absent in the full-length domain. We conclude that to function as a stator in the Tol-Pal complex dimeric TolR must undergo large scale structural remodeling reminiscent of that proposed for MotB, where the N- and C-terminal sequences unfold in order for the protein to both reach and bind the PG layer ∼90 Å away from the inner membrane.
- From the Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU and.
Organizational Affiliation: