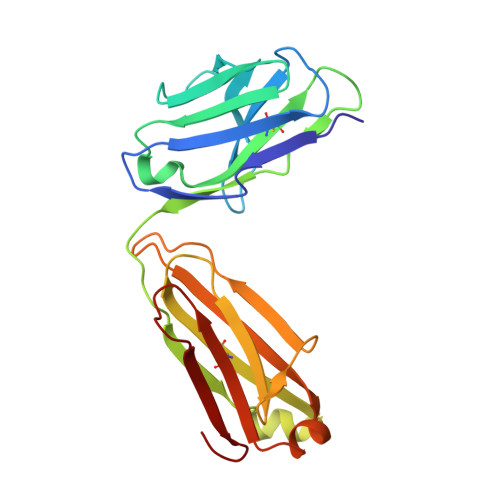
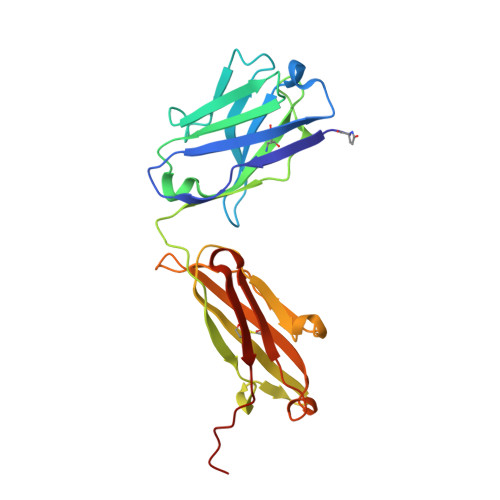
The molecular mode of action and species specificity of canakinumab, a human monoclonal antibody neutralizing IL-1 beta.
Rondeau, J.M., Ramage, P., Zurini, M., Gram, H.(2015) MAbs 7: 1151-1160
- PubMed: 26284424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2015.1081323
- Primary Citation Related Structures:
5BVJ, 5BVP - PubMed Abstract:
Interleukin-1β (IL-1β) plays a key role in autoinflammatory diseases, such as systemic juvenile idiopathic arthritis (sJIA) or cryopyrin-associated periodic syndrome (CAPS). Canakinumab, a human monoclonal anti-IL-1β antibody, was recently approved for human use under the brand name Ilaris®. Canakinumab does not cross-react with IL-1β from mouse, rat, rabbit, or macaques. The crystal structure of the canakinumab Fab bound to human IL-1β was determined in an attempt to rationalize the species specificity. The X-ray analysis reveals a complex surface epitope with an intricate network of well-ordered water molecules at the antibody-antigen interface. The canakinumab paratope is largely pre-organized, as demonstrated by the structure determination of the free Fab. Glu 64 of human IL-1β is a pivotal epitope residue explaining the exquisite species specificity of canakinumab. We identified marmoset as the only non-human primate species that carries Glu 64 in its IL-1β and demonstrates full cross-reactivity of canakinumab, thereby enabling toxicological studies in this species. As demonstrated by the X-ray structure of the complex with IL-1β, canakinumab binds IL-1β on the opposite side with respect to the IL-1RAcP binding site, and in an approximately orthogonal orientation with respect to IL-1RI. However, the antibody and IL-1RI binding sites slightly overlap and the VH region of canakinumab would sterically interfere with the D1 domain of IL-1RI, as shown by a structural overlay with the IL-1β:IL-1RI complex. Therefore, direct competition with IL-1RI for IL-1β binding is the molecular mechanism of neutralization by canakinumab, which is also confirmed by competition assays with recombinant IL-1RI and IL-1RII.
- a Novartis Institutes for BioMedical Research ; Basel , Switzerland.
Organizational Affiliation: