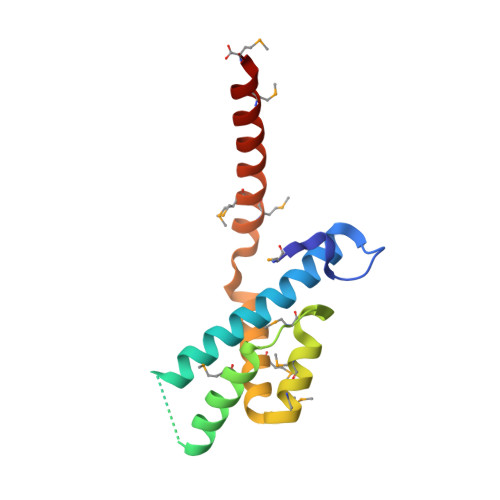
Structural Analysis of the Rubisco-Assembly Chaperone RbcX-II from Chlamydomonas reinhardtii.
Bracher, A., Hauser, T., Liu, C., Hartl, F.U., Hayer-Hartl, M.(2015) PLoS One 10: e0135448-e0135448
- PubMed: 26305355 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0135448
- Primary Citation Related Structures:
5BS1, 5BS2 - PubMed Abstract:
The most prevalent form of the Rubisco enzyme is a complex of eight catalytic large subunits (RbcL) and eight regulatory small subunits (RbcS). Rubisco biogenesis depends on the assistance by specific molecular chaperones. The assembly chaperone RbcX stabilizes the RbcL subunits after folding by chaperonin and mediates their assembly to the RbcL8 core complex, from which RbcX is displaced by RbcS to form active holoenzyme. Two isoforms of RbcX are found in eukaryotes, RbcX-I, which is more closely related to cyanobacterial RbcX, and the more distant RbcX-II. The green algae Chlamydomonas reinhardtii contains only RbcX-II isoforms, CrRbcX-IIa and CrRbcX-IIb. Here we solved the crystal structure of CrRbcX-IIa and show that it forms an arc-shaped dimer with a central hydrophobic cleft for binding the C-terminal sequence of RbcL. Like other RbcX proteins, CrRbcX-IIa supports the assembly of cyanobacterial Rubisco in vitro, albeit with reduced activity relative to cyanobacterial RbcX-I. Structural analysis of a fusion protein of CrRbcX-IIa and the C-terminal peptide of RbcL suggests that the peptide binding mode of RbcX-II may differ from that of cyanobacterial RbcX. RbcX homologs appear to have adapted to their cognate Rubisco clients as a result of co-evolution.
- Department of Cellular Biochemistry, Max-Planck-Institute of Biochemistry, Martinsried, Germany.
Organizational Affiliation: