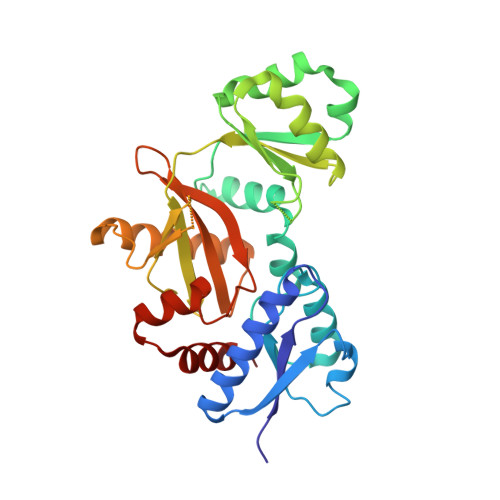
Structure of D-alanine-D-alanine ligase from Yersinia pestis: nucleotide phosphate recognition by the serine loop.
Tran, H.T., Hong, M.K., Ngo, H.P., Huynh, K.H., Ahn, Y.J., Wang, Z., Kang, L.W.(2016) Acta Crystallogr D Struct Biol 72: 12-21
- PubMed: 26894530 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798315021671
- Primary Citation Related Structures:
4ZQI, 5BPF, 5BPH, 5C1O, 5C1P - PubMed Abstract:
D-Alanyl-D-alanine is an essential precursor of bacterial peptidoglycan and is synthesized by D-alanine-D-alanine ligase (DDL) with hydrolysis of ATP; this reaction makes DDL an important drug target for the development of antibacterial agents. Five crystal structures of DDL from Yersinia pestis (YpDDL) were determined at 1.7-2.5 Å resolution: apo, AMP-bound, ADP-bound, adenosine 5'-(β,γ-imido)triphosphate-bound, and D-alanyl-D-alanine- and ADP-bound structures. YpDDL consists of three domains, in which four loops, loop 1, loop 2 (the serine loop), loop 3 (the ω-loop) and loop 4, constitute the binding sites for two D-alanine molecules and one ATP molecule. Some of them, especially the serine loop and the ω-loop, show flexible conformations, and the serine loop is mainly responsible for the conformational change in substrate nucleotide phosphates. Enzyme-kinetics assays were carried out for both the D-alanine and ATP substrates and a substrate-binding mechanism was proposed for YpDDL involving conformational changes of the loops.
- Department of Biological Sciences, Konkuk University, Hwayang-dong, Gwangjin-gu, Seoul 143-701, Republic of Korea.
Organizational Affiliation: