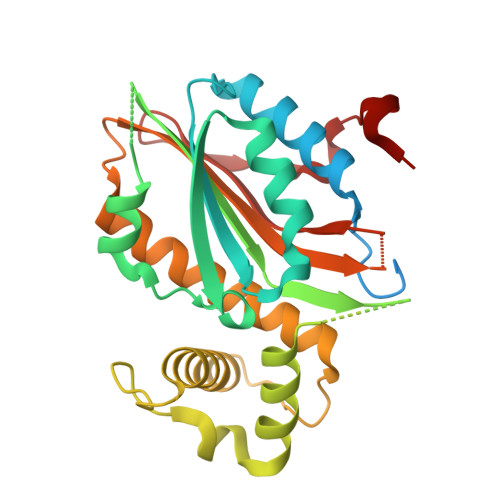
Structural insights into the LCIB protein family reveals a new group of beta-carbonic anhydrases
Jin, S., Sun, J., Wunder, T., Tang, D., Cousins, A.B., Sze, S.K., Mueller-Cajar, O., Gao, Y.G.(2016) Proc Natl Acad Sci U S A 113: 14716-14721
- PubMed: 27911826 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1616294113
- Primary Citation Related Structures:
5B5X, 5B5Y, 5B5Z, 5B60, 5K5W - PubMed Abstract:
Aquatic microalgae have evolved diverse CO 2 -concentrating mechanisms (CCMs) to saturate the carboxylase with its substrate, to compensate for the slow kinetics and competing oxygenation reaction of the key photosynthetic CO 2 -fixing enzyme rubisco. The limiting CO 2 -inducible B protein (LCIB) is known to be essential for CCM function in Chlamydomonas reinhardtii To assign a function to this previously uncharacterized protein family, we purified and characterized a phylogenetically diverse set of LCIB homologs. Three of the six homologs are functional carbonic anhydrases (CAs). We determined the crystal structures of LCIB and limiting CO 2 -inducible C protein (LCIC) from C. reinhardtii and a CA-functional homolog from Phaeodactylum tricornutum, all of which harbor motifs bearing close resemblance to the active site of canonical β-CAs. Our results identify the LCIB family as a previously unidentified group of β-CAs, and provide a biochemical foundation for their function in the microalgal CCMs.
- School of Biological Sciences, Nanyang Technological University, Singapore 637551.
Organizational Affiliation: