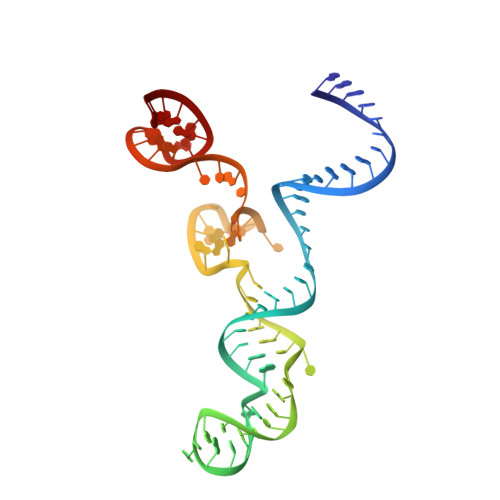
Structural Basis for the Altered PAM Specificities of Engineered CRISPR-Cas9
Hirano, S., Nishimasu, H., Ishitani, R., Nureki, O.(2016) Mol Cell 61: 886-894
- PubMed: 26990991 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2016.02.018
- Primary Citation Related Structures:
5B2R, 5B2S, 5B2T - PubMed Abstract:
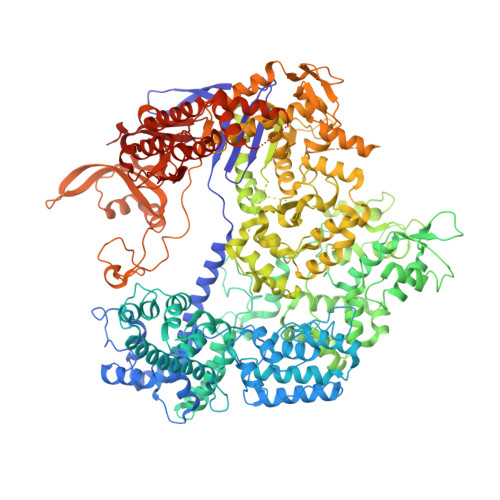
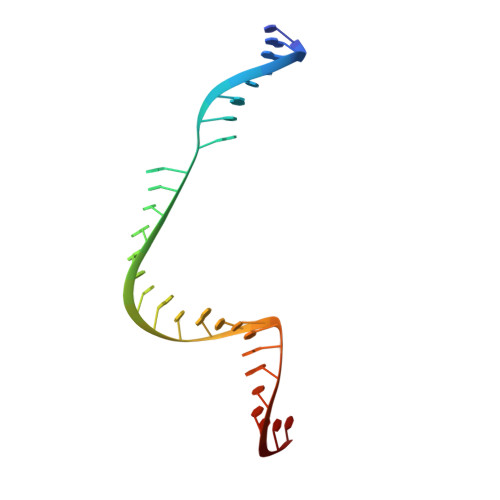
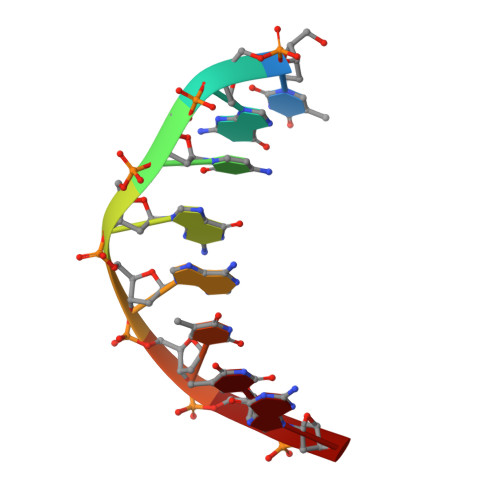
The RNA-guided endonuclease Cas9 cleaves double-stranded DNA targets bearing a PAM (protospacer adjacent motif) and complementarity to the guide RNA. A recent study showed that, whereas wild-type Streptococcus pyogenes Cas9 (SpCas9) recognizes the 5'-NGG-3' PAM, the engineered VQR, EQR, and VRER SpCas9 variants recognize the 5'-NGA-3', 5'-NGAG-3', and 5'-NGCG-3' PAMs, respectively, thus expanding the targetable sequences in Cas9-mediated genome editing applications. Here, we present the high-resolution crystal structures of the three SpCas9 variants in complexes with a single-guide RNA and its altered PAM-containing, partially double-stranded DNA targets. A structural comparison of the three SpCas9 variants with wild-type SpCas9 revealed that the multiple mutations synergistically induce an unexpected displacement in the phosphodiester backbone of the PAM duplex, thereby allowing the SpCas9 variants to directly recognize the altered PAM nucleotides. Our findings explain the altered PAM specificities of the SpCas9 variants and establish a framework for further rational engineering of CRISPR-Cas9.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan.
Organizational Affiliation: