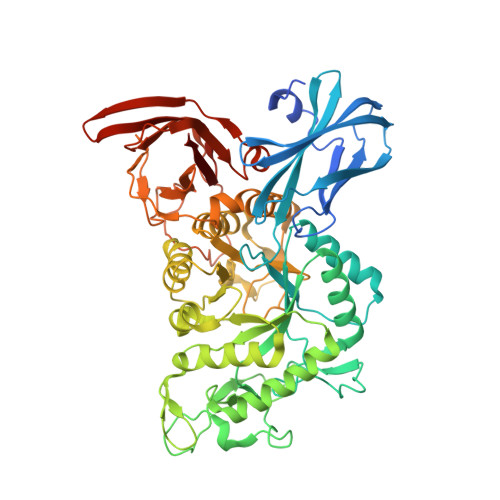
Crystal structure of thermophilic dextranase from Thermoanaerobacter pseudethanolicus
Suzuki, N., Kishine, N., Fujimoto, Z., Sakurai, M., Momma, M., Ko, J.A., Nam, S.H., Kimura, A., Kim, Y.M.(2016) J Biochem 159: 331-339
- PubMed: 26494689 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvv104
- Primary Citation Related Structures:
5AXG, 5AXH - PubMed Abstract:
The crystal structures of the wild type and catalytic mutant Asp-312→Gly in complex with isomaltohexaose of endo-1,6-dextranase from the thermophilic bacterium Thermoanaerobacter pseudethanolicus (TpDex), belonging to the glycoside hydrolase family 66, were determined. TpDex consists of three structural domains, a catalytic domain comprising an (β/α)8-barrel and two β-domains located at both N- and C-terminal ends. The isomaltohexaose-complex structure demonstrated that the isomaltohexaose molecule was bound across the catalytic site, showing that TpDex had six subsites (-4 to +2) in the catalytic cleft. Marked movement of the Trp-376 side-chain along with loop 6, which was the side wall component of the cleft at subsite +1, was observed to occupy subsite +1, indicating that it might expel the cleaved aglycone subsite after the hydrolysis reaction. Structural comparison with other mesophilic enzymes indicated that several structural features of TpDex, loop deletion, salt bridge and surface-exposed charged residue, may contribute to thermostability.
- Biomolecular Research Unit, National Institute of Agrobiological Sciences, 2-1-2 Kannondai, Tsukuba 305-8602, Japan; zui@affrc.go.jp u9897854@jnu.ac.kr.
Organizational Affiliation: