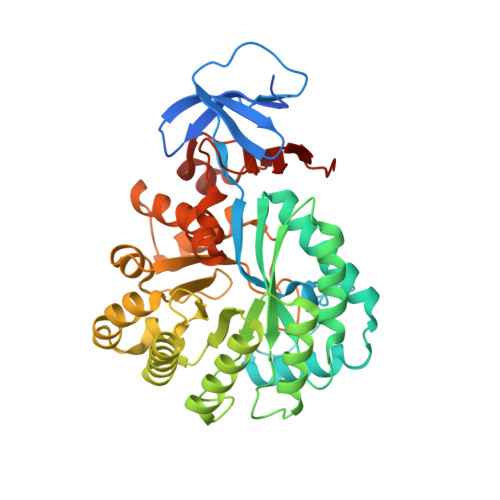
X-Ray Structure and Mutagenesis Studies of the N-Isopropylammelide Isopropylaminohydrolase, Atzc
Balotra, S., Warden, A.C., Newman, J., Briggs, L.J., Scott, C., Peat, T.S.(2015) PLoS One 1: 37700
- PubMed: 26390431 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0137700
- Primary Citation Related Structures:
4CQB, 4CQC, 4CQD, 5AKQ - PubMed Abstract:
The N-isopropylammelide isopropylaminohydrolase from Pseudomonas sp. strain ADP, AtzC, provides the third hydrolytic step in the mineralization of s-triazine herbicides, such as atrazine. We obtained the X-ray crystal structure of AtzC at 1.84 Å with a weak inhibitor bound in the active site and then used a combination of in silico docking and site-directed mutagenesis to understand the interactions between AtzC and its substrate, isopropylammelide. The substitution of an active site histidine residue (His249) for an alanine abolished the enzyme's catalytic activity. We propose a plausible catalytic mechanism, consistent with the biochemical and crystallographic data obtained that is similar to that found in carbonic anhydrase and other members of subtype III of the amidohydrolase family.
- CSIRO Land and Water Flagship, Black Mountain, Canberra, Australia; Research School of Chemistry, Australian National University, Canberra, Australian Capital Territory, Australia.
Organizational Affiliation: